Understanding genetic variation in cancer using targeted long-read sequencing
- shared.published_on: September 26 2019
- shared.source: Jackson Lab Long-Read Sequencing Workshop
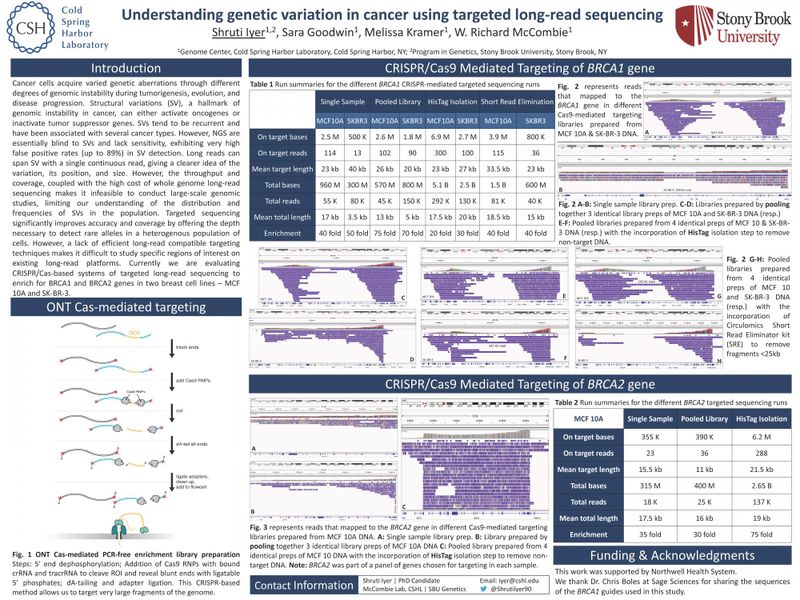
Cancer cells acquire varied genetic aberrations through different degrees of genomic instability during tumorigenesis, evolution, and disease progression. Structural variations (SV), a hallmark of genomic instability in cancer, can either activate oncogenes or inactivate tumor suppressor genes. SVs tend to be recurrent and have been associated with several cancer types. However, NGS are essentially blind to SVs and lack sensitivity, exhibiting very high false positive rates (up to 89%) in SV detection. Long reads can span SV with a single continuous read, giving a clearer idea of the variation, its position, and size. However, the throughput and coverage, coupled with the high cost of whole genome long-read sequencing makes it infeasible to conduct large-scale genomic studies, limiting our understanding of the distribution and frequencies of SVs in the population. Targeted sequencing significantly improves accuracy and coverage by offering the depth necessary to detect rare alleles in a heterogenous population of cells. However, a lack of efficient long-read compatible targeting techniques makes it difficult to study specific regions of interest on existing long-read platforms. Currently we are evaluating CRISPR/Cas-based systems of targeted long-read sequencing to enrich for BRCA1 and BRCA2 genes in two breast cell lines – MCF 10A and SK-BR-3.






