Influenza virus sequencing from RNA using SQK-NBD114 (.24 or .96) (INF_9189_v114_revN_07Oct2025)
MinION: Protocol
V INF_9189_v114_revN_07Oct2025
FOR RESEARCH USE ONLY
Contents
Introduction to the protocol
Library preparation
- 3. Reverse transcription, PCR and clean-up
- 4. End-prep
- 5. Native barcode ligation
- 6. Adapter ligation and clean-up
- 7. Priming and loading the MinION and GridION Flow Cell
Sequencing and data analysis
Troubleshooting
1. Overview of the protocol
Introduction to the influenza whole genome sequencing protocol
This protocol has been upgraded to our Kit 14 chemistry and describes how to carry out PCR amplification and native barcoding of influenza amplicons using the Native Barcoding Kit 24 or 96 V14 (SQK-NBD114.24 or SQK-NBD114.96). There are 96 unique barcodes available, allowing the user to pool up to 96 different Influenza A and/or Influenza B samples in one sequencing experiment.
While this protocol is available in the Nanopore Community, we kindly ask users to ensure they are citing the following references, that this protocol is based on.
Single-reaction genomic amplification accelerates sequencing and vaccine production for classical and Swine origin human influenza A viruses by Bin Zhou *et al*., 2009. and Universal influenza B virus genomic amplification facilitates sequencing, diagnostics, and reverse genetics by Bin Zhou *et al*., 2014.
Steps in the sequencing workflow:
Prepare for your experiment
You will need to:
- Ensure you have the sequencing kit, the correct equipment and third-party reagents.
- Download the software for acquiring and analysing your data.
- Check your flow cell to ensure it has enough pores for a good sequencing run.
Prepare your library
You will need to:
- Perform RT-PCR amplification on influenza samples.
- Prepare the DNA ends for adapter attachment.
- Ligate native barcodes supplied in the kit to the DNA ends.
- Ligate sequencing adapters supplied in the kit to the DNA ends.
- Prime the flow cell, and load your DNA library into the flow cell.
Sequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software, which will collect raw data from the device and basecall the reads.
- Demultiplex barcoded reads in MinKNOW choosing the SQK-NBD114.24 or SQK-NBD114.96 kit option.
- Analyse your data using the Influenza typing workflow (wf-flu) in EPI2ME Labs.
We do not recommend mixing barcoded libraries with non-barcoded libraries prior to sequencing.
Compatibility of this protocol
This protocol should only be used in combination with:
- Native Barcoding Kit 24 V14 (SQK-NBD114.24)
- Native Barcoding Kit 96 V14 (SQK-NBD114.96)
- R10.4.1 flow cells (FLO-MIN114)
- Flow Cell Wash Kit (EXP-WSH004)
- Sequencing Auxiliary Vials V14 (EXP-AUX003)
- Native Barcoding Expansion V14 (EXP-NBA114)
- MinION Mk1D - MinION Mk1D IT requirements document
- GridION - GridION IT requirements document
2. Equipment and consumables
材料
- Input influenza RNA
- Influenza A primers
- Influenza B primers
- Native Barcoding Kit 24 V14 (SQK-NBD114.24) OR Native Barcoding Kit 96 V14 (SQK-NBD114.96)
- SFB Expansion (EXP-SFB001)
耗材
- SuperScript III One-Step RT-PCR System with Platinum Taq DNA Polymerase (ThermoFisher, cat # 12574018 or 12574026)
- 无核酸酶水(如 Thermo Scientific,AM9937)
- Agencourt AMPure XP 磁珠(Beckman Coulter™, A63881)
- 新制备的 80% 乙醇(用无核酸酶水配制)
- NEB Blunt/TA 连接酶预混液(NEB,M0367)
- NEBNext Ultra II 末端修复/ dA尾添加模块(NEB,E7546)
- NEBNext 快速连接模块(NEB,E6056)
- Eppendorf 低吸附 twin.tec® 96 孔 PCR 板,半裙边(Merck,EP0030129504)带热封
- 1.5 ml Eppendorf DNA LoBind 离心管
- 2 ml Eppendorf DNA LoBind 离心管
- 5 ml Eppendorf DNA LoBind 离心管
- 15 ml Eppendorf DNA LoBind tubes
- Reagent reservoirs for multichannel pipetting
- Qubit™ 分析管(Invitrogen, Q32856)
- Qubit™ dsDNA HS Assay(双链DNA高灵敏度检测)试剂盒(ThermoFisher,Q32851)
- 牛血清白蛋白(BSA)(50 mg/mL)(例如 Invitrogen™ UltraPure™ BSA 50 mg/mL, AM2616)
仪器
- 适用于1.5ml Eppendorf 离心管的磁力架
- Magnetic rack suitable for 96 well plates, e.g. DynaMag™-96 Side Skirted Magnet (Thermo Fisher CAT#12027)
- 迷你离心机
- 涡旋混匀仪
- 热循环仪
- Hula混匀仪(低速旋转式混匀仪)
- 微孔板离心机
- 多通道移液枪和枪头
- P1000 移液枪和枪头
- P200 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P10 移液枪和枪头
- P2 移液枪和枪头
- 盛有冰的冰桶
- 计时器
- Qubit™ 荧光计(或用于质控检测的等效仪器)
可选仪器
- PCR hoods with UV steriliser
- PCR-Cooler (Eppendorf)
- Eppendorf 5424 离心机(或等效器材)
For this protocol, you will need your extracted Influenza RNA input in 10 mM Tris-HCl, pH 8.0.
A minimum volume of 1 µl is required per sample input of Influenza A or B.
If performing typing of an unknown sample, 2 µl of input will be required:
- 1 µl input for the Influenza A primer mix
- 1 µl for the Influenza B primer mix
Before starting
This protocol outlines how to carry out PCR amplification and native barcoding of influenza amplicons from multiple samples on a 96-well plate using the Native Barcoding Kit 24 or 96 V14 (SQK-NBD114.24 or SQK-NBD114.96).
When processing multiple samples simultaneously, we recommend making master mixes with an additional 10% of the volume. We also recommend using a template-free pre-PCR hood for making up the master mixes, and a separate template pre-PCR hood for handling the samples. It is important to clean and/or UV irradiate these hoods between sample batches. Furthermore, to track and monitor cross-contamination events, it is important to run a negative control reaction at the reverse transcription stage using nuclease-free water instead of sample, and carrying this control through the rest of the prep.
All post-PCR procedures must be carried out in a separate area to the pre-PCR preparation, with dedicated equipment for liquid handling in each area.
Check your flow cell
We highly recommend that you check the number of pores in your flow cell prior to starting a sequencing experiment. This should be done within 12 weeks of purchasing your MinION/GridION Flow Cells. Oxford Nanopore Technologies will replace any unused flow cell with fewer than the number of pores listed in the Table below, when the result is reported within two days of performing the flow cell check, and when the storage recommendations have been followed. To do the flow cell check, please follow the instructions in the Flow Cell Check document.
| Flow cell | Minimum number of active pores covered by warranty |
|---|---|
| MinION/GridION Flow Cell | 800 |
Third-party reagents
We have validated and recommend the use of all the third-party reagents used in this protocol. Alternatives have not been tested by Oxford Nanopore Technologies.
For all third-party reagents, we recommend following the manufacturer's instructions to prepare the reagents for use.
Influenza primer sequences
Influenza A primer sequences described in the protocol originated from: Single-reaction genomic amplification accelerates sequencing and vaccine production for classical and Swine origin human influenza A viruses by Bin Zhou *et al*., 2009.
| Component | Sequence |
|---|---|
| Tuni 12 | ACGCGTGATCAGCAAAAGCAGG |
| Tuni 12.4 | ACGCGTGATCAGCGAAAGCAGG |
| Tuni 13 | ACGCGTGATCAGTAGAAACAAGG |
Influenza B primer sequences described in the protocol originated from: Universal influenza B virus genomic amplification facilitates sequencing, diagnostics, and reverse genetics by Bin Zhou *et al*., 2014.
| Component | Sequence |
|---|---|
| B-PBs-UniF | GGGGGGAGCAGAAGCGGAGC |
| B-PBs-UniR | CCGGGTTATTAGTAGAAACACGAGC |
| B-PA-UniF | GGGGGGAGCAGAAGCGGTGC |
| B-PA-UniR | CCGGGTTATTAGTAGAAACACGTGC |
| B-HANA-UniF | GGGGGGAGCAGAAGCAGAGC |
| B-HANA-UniR | CCGGGTTATTAGTAGTAACAAGAGC |
| B-NP-UniF | GGGGGGAGCAGAAGCACAGC |
| B-NP-UniR | CCGGGTTATTAGTAGAAACAACAGC |
| B-M-Uni3F | GGGGGGAGCAGAAGCACGCACTT |
| B-Mg-Uni3F | GGGGGGAGCAGAAGCAGGCACTT |
| B-M-Uni3R | CCGGGTTATTAGTAGAAACAACGCACTT |
| B-NS-Uni3F | GGGGGGAGCAGAAGCAGAGGATT |
| B-NS-Uni3R | CCGGGTTATTAGTAGTAACAAGAGGATT |
Short Fragment Buffer (SFB)
Within the Native Barcoding Kit 24 V14 (SQK-NBD114.24) and Native Barcoding Kit 96 V14 (SQK-NBD114.96), Short Fragment Buffer (SFB) is supplied at the volume needed to complete the "Reverse transcription, PCR and clean-up" and "Adapter ligation and clean-up" steps of the protocol.
However, extra Short Fragment Buffer (SFB) is required for the "native barcode ligation" step of the protocol. This can be purchased with our SFB Expansion (EXP-SFB001).
AMPure XP Beads
Within the Native Barcoding Kits V14 (SQK-NBD114.24 and SQK-NBD114.96), AMPure XP Beads (AXP) are supplied at the volume needed to complete the native barcoding sections of the protocol: "Native barcode ligation" and "Adapter ligation and clean-up".
However, extra AMPure XP Beads are required for the "Reverse transcription, PCR and clean-up" step of the protocol.
The Native Adapter (NA) used in this kit and protocol is not interchangeable with other sequencing adapters.
免扩增条形码测序试剂盒-24 V14(SQK-NBD114.24)内容物
请注意: 我们正在更新免扩增条形码建库试剂盒,新版将增加短片段缓冲液(SFB)的体积。如果您使用的是旧版试剂盒,或需要额外的短片段缓冲液(SFB),可通过购买 SFB 扩展包(EXP-SFB001) 获取。
新版试剂盒:短片段缓冲液体积增加(SFB)
| 名称 | 缩写 | 管盖颜色 | 管数 | 每管溶液体积 (μl) |
|---|---|---|---|---|
| 免扩增条形码 | NB01-24 | - | 两板,每板三套条形码组合 | 每孔 5 µl |
| DNA 参照 | DCS | 黄色 | 2 | 35 |
| 免扩增接头 | NA | 绿色 | 1 | 40 |
| 测序缓冲液 | SB | 红色 | 1 | 700 |
| 文库颗粒 | LIB | 粉色 | 1 | 600 |
| 文库溶液 | LIS | 白色管盖,粉色标签 | 1 | 600 |
| 洗脱缓冲液 | EB | 黑色 | 2 | 500 |
| AMPure XP 磁珠 | AXP | 透明管盖,浅青绿色标签 | 1 | 6000 |
| 长片段缓冲液 | LFB | 橙色 | 1 | 1800 |
| 短片段缓冲液 | SFB | 透明 | 1 | 13000 |
| EDTA | EDTA | 蓝色 | 1 | 700 |
| 测序芯片冲洗液 | FCF | 透明管盖,浅蓝色标签 | 1 | 8000 |
| 测序芯片系绳 | FCT | 紫色 | 1 | 200 |
旧版试剂盒:较低体积的短片段缓冲液(SFB)
| 名称 | 缩写 | 管盖颜色 | 管数 | 每管溶液体积 (μl) |
|---|---|---|---|---|
| 免扩增条形码 | NB01-24 | - | 两板,每板三套条形码组合 | 每孔 5 µl |
| DNA 参照 | DCS | 黄色 | 2 | 35 |
| 免扩增接头 | NA | 绿色 | 1 | 40 |
| 测序缓冲液 | SB | 红色 | 1 | 700 |
| 文库颗粒 | LIB | 粉色 | 1 | 600 |
| 文库溶液 | LIS | 白色管盖,粉色标签 | 1 | 600 |
| 洗脱缓冲液 | EB | 黑色 | 2 | 500 |
| AMPure XP 磁珠 | AXP | 透明管盖,浅青绿色标签 | 1 | 6000 |
| 长片段缓冲液 | LFB | 橙色 | 1 | 1800 |
| 短片段缓冲液 | SFB | 透明 | 1 | 1800 |
| EDTA | EDTA | 透明 | 1 | 700 |
| 测序芯片冲洗液 | FCF | 透明管盖,浅蓝色标签 | 1 | 8000 |
| 测序芯片系绳 | FCT | 紫色 | 1 | 200 |
请注意: 本产品包含由贝克曼库尔特公司(Beckman Coulter, Inc)生产的 AMPure XP 试剂,并可与试剂盒一起于-20℃下储存(试剂稳定性将不受损害)。
请注意: DNA参照(DCS)是一段可比对到Lambda基因组的3'端、长度为3.6 kb 的标准扩增子。
免扩增条形码测序试剂盒-96 V14(SQK-NBD114.96)内容物
请注意: 我们正在更新免扩增条形码建库试剂盒,新版将增加短片段缓冲液(SFB)的体积。如果您使用的是旧版试剂盒,或需要额外的短片段缓冲液(SFB),可通过购买 SFB 扩展包(EXP-SFB001) 获取。
新版试剂盒:短片段缓冲液体积增加(SFB)
| 名称 | 缩写 | 管盖颜色 | 管数 | 每管溶液体积 (μl) |
|---|---|---|---|---|
| 免扩增条形码 | NB01-96 | - | 3 盘 | 每孔 8 μl |
| DNA 参照 | DCS | 黄色 | 3 | 35 |
| 免扩增接头 | NA | 绿色 | 2 | 40 |
| 测序缓冲液 | SB | 红色 | 2 | 700 |
| 文库颗粒 | LIB | 粉色 | 2 | 600 |
| 文库溶液 | LIS | 白色管盖,粉色标签 | 2 | 600 |
| 洗脱缓冲液 | EB | 黑色 | 1 | 1500 |
| AMPure XP 磁珠 | AXP | 琥珀色 | 1 | 6000 |
| 长片段缓冲液 | LFB | 橙色 | 1 | 7500 |
| 短片段缓冲液 | SFB | 透明 | 1 | 25000 |
| 测序芯片冲洗液 | FCF | 蓝色 | 1 | 15500 |
| 测序芯片系绳 | FCT | 紫色 | 2 | 200 |
| EDTA | EDTA | 透明 | 1 | 700 |
旧版试剂盒:较低体积的短片段缓冲液(SFB)
| 名称 | 缩写 | 管盖颜色 | 管数 | 每管溶液体积 (μl) |
|---|---|---|---|---|
| 免扩增条形码 | NB01-96 | - | 3 盘 | 每孔 8 μl |
| DNA 参照 | DCS | 黄色 | 3 | 35 |
| 免扩增接头 | NA | 绿色 | 2 | 40 |
| 测序缓冲液 | SB | 红色 | 2 | 700 |
| 文库颗粒 | LIB | 粉色 | 2 | 600 |
| 文库溶液 | LIS | 白色管盖,粉色标签 | 2 | 600 |
| 洗脱缓冲液 | EB | 黑色 | 1 | 1500 |
| AMPure XP 磁珠 | AXP | 琥珀色 | 1 | 6000 |
| 长片段缓冲液 | LFB | 橙色 | 1 | 7500 |
| 短片段缓冲液 | SFB | 透明 | 1 | 7500 |
| 测序芯片冲洗液 | FCF | 蓝色 | 1 | 15500 |
| 测序芯片系绳 | FCT | 紫色 | 2 | 200 |
| EDTA | EDTA | 透明 | 1 | 700 |
请注意: 本产品包含由贝克曼库尔特公司(Beckman Coulter, Inc)生产的 AMPure XP 试剂,并可与试剂盒一起于-20℃下储存(试剂稳定性将不受损害)。
条形码孔板中的条形码是按列排序的。
请注意: DNA参照(DCS)是一段可比对到Lambda基因组的3'端、长度为3.6 kb 的标准扩增子。
To maximise the use of the Native Barcoding Kits, the Native Barcode Auxiliary V14 (EXP-NBA114) and the Sequencing Auxiliary Vials V14 (EXP-AUX003) expansion packs are available.
These expansions provide extra library preparation and flow cell priming reagents to allow users to utilise any unused barcodes for those running in smaller subsets.
Both expansion packs used together will provide enough reagents for 12 reactions. For customers requiring extra EDTA to maximise the use of barcodes, we recommend using 0.25 M EDTA and adding 4 µl for library preps using the SQK-NBD114.24 kit and 2 µl for preps using the SQK-NBD114.96 kit.
Native Barcode Auxiliary V14 (EXP-NBA114) contents:
Note: This product contains AMPure XP reagent manufactured by Beckman Coulter, Inc. and can be stored at -20°C with the kit without detriment to reagent stability.
Sequencing Auxiliary Vials V14 (EXP-AUX003) contents:
Native barcode sequences
Below is the full list of our native barcode (NB01-96) sequences. The first 24 unique barcodes are available in the Native Barcoding Kit 24 V14 (SQK-NBD114.24). The Native Barcoding Kit 96 V14 (SQK-NBD114.96) include the first 24 native barcodes, with the additional 72 unique barcodes. The native barcodes are shipped at 640 nM.
In addition to the barcodes, there are also flanking sequences which add an extra level of context during analysis.
Barcode flanking sequences:
Forward sequence: 5' - AAGGTTAA - barcode - CAGCACCT - 3' Reverse sequence: 5' - GGTGCTG - barcode - TTAACCTTAGCAAT - 3'
Native barcode sequences
| Component | Forward sequence | Reverse sequence |
|---|---|---|
| NB01 | CACAAAGACACCGACAACTTTCTT | AAGAAAGTTGTCGGTGTCTTTGTG |
| NB02 | ACAGACGACTACAAACGGAATCGA | TCGATTCCGTTTGTAGTCGTCTGT |
| NB03 | CCTGGTAACTGGGACACAAGACTC | GAGTCTTGTGTCCCAGTTACCAGG |
| NB04 | TAGGGAAACACGATAGAATCCGAA | TTCGGATTCTATCGTGTTTCCCTA |
| NB05 | AAGGTTACACAAACCCTGGACAAG | CTTGTCCAGGGTTTGTGTAACCTT |
| NB06 | GACTACTTTCTGCCTTTGCGAGAA | TTCTCGCAAAGGCAGAAAGTAGTC |
| NB07 | AAGGATTCATTCCCACGGTAACAC | GTGTTACCGTGGGAATGAATCCTT |
| NB08 | ACGTAACTTGGTTTGTTCCCTGAA | TTCAGGGAACAAACCAAGTTACGT |
| NB09 | AACCAAGACTCGCTGTGCCTAGTT | AACTAGGCACAGCGAGTCTTGGTT |
| NB10 | GAGAGGACAAAGGTTTCAACGCTT | AAGCGTTGAAACCTTTGTCCTCTC |
| NB11 | TCCATTCCCTCCGATAGATGAAAC | GTTTCATCTATCGGAGGGAATGGA |
| NB12 | TCCGATTCTGCTTCTTTCTACCTG | CAGGTAGAAAGAAGCAGAATCGGA |
| NB13 | AGAACGACTTCCATACTCGTGTGA | TCACACGAGTATGGAAGTCGTTCT |
| NB14 | AACGAGTCTCTTGGGACCCATAGA | TCTATGGGTCCCAAGAGACTCGTT |
| NB15 | AGGTCTACCTCGCTAACACCACTG | CAGTGGTGTTAGCGAGGTAGACCT |
| NB16 | CGTCAACTGACAGTGGTTCGTACT | AGTACGAACCACTGTCAGTTGACG |
| NB17 | ACCCTCCAGGAAAGTACCTCTGAT | ATCAGAGGTACTTTCCTGGAGGGT |
| NB18 | CCAAACCCAACAACCTAGATAGGC | GCCTATCTAGGTTGTTGGGTTTGG |
| NB19 | GTTCCTCGTGCAGTGTCAAGAGAT | ATCTCTTGACACTGCACGAGGAAC |
| NB20 | TTGCGTCCTGTTACGAGAACTCAT | ATGAGTTCTCGTAACAGGACGCAA |
| NB21 | GAGCCTCTCATTGTCCGTTCTCTA | TAGAGAACGGACAATGAGAGGCTC |
| NB22 | ACCACTGCCATGTATCAAAGTACG | CGTACTTTGATACATGGCAGTGGT |
| NB23 | CTTACTACCCAGTGAACCTCCTCG | CGAGGAGGTTCACTGGGTAGTAAG |
| NB24 | GCATAGTTCTGCATGATGGGTTAG | CTAACCCATCATGCAGAACTATGC |
| NB25 | GTAAGTTGGGTATGCAACGCAATG | CATTGCGTTGCATACCCAACTTAC |
| NB26 | CATACAGCGACTACGCATTCTCAT | ATGAGAATGCGTAGTCGCTGTATG |
| NB27 | CGACGGTTAGATTCACCTCTTACA | TGTAAGAGGTGAATCTAACCGTCG |
| NB28 | TGAAACCTAAGAAGGCACCGTATC | GATACGGTGCCTTCTTAGGTTTCA |
| NB29 | CTAGACACCTTGGGTTGACAGACC | GGTCTGTCAACCCAAGGTGTCTAG |
| NB30 | TCAGTGAGGATCTACTTCGACCCA | TGGGTCGAAGTAGATCCTCACTGA |
| NB31 | TGCGTACAGCAATCAGTTACATTG | CAATGTAACTGATTGCTGTACGCA |
| NB32 | CCAGTAGAAGTCCGACAACGTCAT | ATGACGTTGTCGGACTTCTACTGG |
| NB33 | CAGACTTGGTACGGTTGGGTAACT | AGTTACCCAACCGTACCAAGTCTG |
| NB34 | GGACGAAGAACTCAAGTCAAAGGC | GCCTTTGACTTGAGTTCTTCGTCC |
| NB35 | CTACTTACGAAGCTGAGGGACTGC | GCAGTCCCTCAGCTTCGTAAGTAG |
| NB36 | ATGTCCCAGTTAGAGGAGGAAACA | TGTTTCCTCCTCTAACTGGGACAT |
| NB37 | GCTTGCGATTGATGCTTAGTATCA | TGATACTAAGCATCAATCGCAAGC |
| NB38 | ACCACAGGAGGACGATACAGAGAA | TTCTCTGTATCGTCCTCCTGTGGT |
| NB39 | CCACAGTGTCAACTAGAGCCTCTC | GAGAGGCTCTAGTTGACACTGTGG |
| NB40 | TAGTTTGGATGACCAAGGATAGCC | GGCTATCCTTGGTCATCCAAACTA |
| NB41 | GGAGTTCGTCCAGAGAAGTACACG | CGTGTACTTCTCTGGACGAACTCC |
| NB42 | CTACGTGTAAGGCATACCTGCCAG | CTGGCAGGTATGCCTTACACGTAG |
| NB43 | CTTTCGTTGTTGACTCGACGGTAG | CTACCGTCGAGTCAACAACGAAAG |
| NB44 | AGTAGAAAGGGTTCCTTCCCACTC | GAGTGGGAAGGAACCCTTTCTACT |
| NB45 | GATCCAACAGAGATGCCTTCAGTG | CACTGAAGGCATCTCTGTTGGATC |
| NB46 | GCTGTGTTCCACTTCATTCTCCTG | CAGGAGAATGAAGTGGAACACAGC |
| NB47 | GTGCAACTTTCCCACAGGTAGTTC | GAACTACCTGTGGGAAAGTTGCAC |
| NB48 | CATCTGGAACGTGGTACACCTGTA | TACAGGTGTACCACGTTCCAGATG |
| NB49 | ACTGGTGCAGCTTTGAACATCTAG | CTAGATGTTCAAAGCTGCACCAGT |
| NB50 | ATGGACTTTGGTAACTTCCTGCGT | ACGCAGGAAGTTACCAAAGTCCAT |
| NB51 | GTTGAATGAGCCTACTGGGTCCTC | GAGGACCCAGTAGGCTCATTCAAC |
| NB52 | TGAGAGACAAGATTGTTCGTGGAC | GTCCACGAACAATCTTGTCTCTCA |
| NB53 | AGATTCAGACCGTCTCATGCAAAG | CTTTGCATGAGACGGTCTGAATCT |
| NB54 | CAAGAGCTTTGACTAAGGAGCATG | CATGCTCCTTAGTCAAAGCTCTTG |
| NB55 | TGGAAGATGAGACCCTGATCTACG | CGTAGATCAGGGTCTCATCTTCCA |
| NB56 | TCACTACTCAACAGGTGGCATGAA | TTCATGCCACCTGTTGAGTAGTGA |
| NB57 | GCTAGGTCAATCTCCTTCGGAAGT | ACTTCCGAAGGAGATTGACCTAGC |
| NB58 | CAGGTTACTCCTCCGTGAGTCTGA | TCAGACTCACGGAGGAGTAACCTG |
| NB59 | TCAATCAAGAAGGGAAAGCAAGGT | ACCTTGCTTTCCCTTCTTGATTGA |
| NB60 | CATGTTCAACCAAGGCTTCTATGG | CCATAGAAGCCTTGGTTGAACATG |
| NB61 | AGAGGGTACTATGTGCCTCAGCAC | GTGCTGAGGCACATAGTACCCTCT |
| NB62 | CACCCACACTTACTTCAGGACGTA | TACGTCCTGAAGTAAGTGTGGGTG |
| NB63 | TTCTGAAGTTCCTGGGTCTTGAAC | GTTCAAGACCCAGGAACTTCAGAA |
| NB64 | GACAGACACCGTTCATCGACTTTC | GAAAGTCGATGAACGGTGTCTGTC |
| NB65 | TTCTCAGTCTTCCTCCAGACAAGG | CCTTGTCTGGAGGAAGACTGAGAA |
| NB66 | CCGATCCTTGTGGCTTCTAACTTC | GAAGTTAGAAGCCACAAGGATCGG |
| NB67 | GTTTGTCATACTCGTGTGCTCACC | GGTGAGCACACGAGTATGACAAAC |
| NB68 | GAATCTAAGCAAACACGAAGGTGG | CCACCTTCGTGTTTGCTTAGATTC |
| NB69 | TACAGTCCGAGCCTCATGTGATCT | AGATCACATGAGGCTCGGACTGTA |
| NB70 | ACCGAGATCCTACGAATGGAGTGT | ACACTCCATTCGTAGGATCTCGGT |
| NB71 | CCTGGGAGCATCAGGTAGTAACAG | CTGTTACTACCTGATGCTCCCAGG |
| NB72 | TAGCTGACTGTCTTCCATACCGAC | GTCGGTATGGAAGACAGTCAGCTA |
| NB73 | AAGAAACAGGATGACAGAACCCTC | GAGGGTTCTGTCATCCTGTTTCTT |
| NB74 | TACAAGCATCCCAACACTTCCACT | AGTGGAAGTGTTGGGATGCTTGTA |
| NB75 | GACCATTGTGATGAACCCTGTTGT | ACAACAGGGTTCATCACAATGGTC |
| NB76 | ATGCTTGTTACATCAACCCTGGAC | GTCCAGGGTTGATGTAACAAGCAT |
| NB77 | CGACCTGTTTCTCAGGGATACAAC | GTTGTATCCCTGAGAAACAGGTCG |
| NB78 | AACAACCGAACCTTTGAATCAGAA | TTCTGATTCAAAGGTTCGGTTGTT |
| NB79 | TCTCGGAGATAGTTCTCACTGCTG | CAGCAGTGAGAACTATCTCCGAGA |
| NB80 | CGGATGAACATAGGATAGCGATTC | GAATCGCTATCCTATGTTCATCCG |
| NB81 | CCTCATCTTGTGAAGTTGTTTCGG | CCGAAACAACTTCACAAGATGAGG |
| NB82 | ACGGTATGTCGAGTTCCAGGACTA | TAGTCCTGGAACTCGACATACCGT |
| NB83 | TGGCTTGATCTAGGTAAGGTCGAA | TTCGACCTTACCTAGATCAAGCCA |
| NB84 | GTAGTGGACCTAGAACCTGTGCCA | TGGCACAGGTTCTAGGTCCACTAC |
| NB85 | AACGGAGGAGTTAGTTGGATGATC | GATCATCCAACTAACTCCTCCGTT |
| NB86 | AGGTGATCCCAACAAGCGTAAGTA | TACTTACGCTTGTTGGGATCACCT |
| NB87 | TACATGCTCCTGTTGTTAGGGAGG | CCTCCCTAACAACAGGAGCATGTA |
| NB88 | TCTTCTACTACCGATCCGAAGCAG | CTGCTTCGGATCGGTAGTAGAAGA |
| NB89 | ACAGCATCAATGTTTGGCTAGTTG | CAACTAGCCAAACATTGATGCTGT |
| NB90 | GATGTAGAGGGTACGGTTTGAGGC | GCCTCAAACCGTACCCTCTACATC |
| NB91 | GGCTCCATAGGAACTCACGCTACT | AGTAGCGTGAGTTCCTATGGAGCC |
| NB92 | TTGTGAGTGGAAAGATACAGGACC | GGTCCTGTATCTTTCCACTCACAA |
| NB93 | AGTTTCCATCACTTCAGACTTGGG | CCCAAGTCTGAAGTGATGGAAACT |
| NB94 | GATTGTCCTCAAACTGCCACCTAC | GTAGGTGGCAGTTTGAGGACAATC |
| NB95 | CCTGTCTGGAAGAAGAATGGACTT | AAGTCCATTCTTCTTCCAGACAGG |
| NB96 | CTGAACGGTCATAGAGTCCACCAT | ATGGTGGACTCTATGACCGTTCAG |
3. Reverse transcription, PCR and clean-up
材料
- Input influenza RNA
- Influenza A primers
- Influenza B primers
耗材
- SuperScript III One-Step RT-PCR System with Platinum Taq DNA Polymerase (ThermoFisher, cat # 12574018 or 12574026)
- 无核酸酶水(如ThermoFisher,AM9937)
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Cat # 0030129504) with heat seals
- 1.5 ml Eppendorf DNA LoBind 离心管
- 5 ml Eppendorf DNA LoBind 离心管
- 15 ml Eppendorf DNA LoBind tubes
- Agencourt AMPure XP 磁珠(Beckman Coulter,A63881)
- 新制备的 80% 乙醇(用无核酸酶水配制)
- Qubit dsDNA HS Assay(双链DNA高灵敏度检测)试剂盒(Invitrogen, Q32851)
- Qubit™ 分析管(Invitrogen, Q32856)
仪器
- 热循环仪
- Qubit™ 荧光计(或用于质控检测的等效仪器)
- Multichannel pipettes suitable for dispensing 20–200 μl, and tips
- P1000 移液枪和枪头
- P200 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P10 移液枪和枪头
- P2 移液枪和枪头
- Magnetic rack suitable for 96 well plates, e.g. DynaMag™-96 Side Skirted Magnet (Thermo Fisher CAT#12027)
- 微孔板离心机
- 迷你离心机
可选仪器
- PCR hoods with UV steriliser
- PCR-Cooler (Eppendorf) or ice bucket with ice
Keep the RNA sample on ice as much as possible to prevent nucleolytic degradation, which may affect sensitivity.
To reduce risk of contamination, we recommend the use of PCR hoods with a UV steriliser when setting up the PCR plates.
- When handling the primer stocks and derivatives, use a clean template-free PCR hood.
- When handling the samples and/or a positive control, use a clean template-addition PCR hood.
In a clean template-free pre-PCR hood, prepare the primer mixes for influenza A and influenza B as follows in 1.5 ml Eppendorf DNA LoBind tubes:
Note: The volume requirements can be adjusted according to stock concentrations and experiment needs.
Influenza A primer mix
| Primer | Concentration | Volume |
|---|---|---|
| Nuclease-free water | - | 378 µl |
| Tuni 12 | 100 µM | 16.8 µl |
| Tuni 12.4 | 100 µM | 4.2 µl |
| Tuni 13 | 100 µM | 21 µl |
| Total | 420 µl |
Influenza B primer mix
| Primer | Concentration | Volume |
|---|---|---|
| Nuclease-free water | - | 378 µl |
| B-PBs-UniF | 100 µM | 5 µl |
| B-PBs-UniR | 100 µM | 5 µl |
| B-PA-UniF | 100 µM | 2.5 µl |
| B-PA-UniR | 100 µM | 2.5 µl |
| B-HANA-UniF | 100 µM | 5 µl |
| B-HANA-UniR | 100 µM | 5 µl |
| B-NP-UniF | 100 µM | 3 µl |
| B-NP-UniR | 100 µM | 3 µl |
| B-M-Uni3F | 100 µM | 1.5 µl |
| B-Mg-Uni3F | 100 µM | 1.5 µl |
| B-M-Uni3R | 100 µM | 3 µl |
| B-NS-Uni3F | 100 µM | 2.5 µl |
| B-NS-Uni3R | 100 µM | 2.5 µl |
| Total | 420 µl |
In the template-free pre-PCR hood, prepare the following master mixes in Eppendorf DNA LoBind tubes and mix thoroughly as follows:
For X12 samples, use 1.5 ml Eppendorf DNA LoBind tubes:
| Component | Influenza A RT-PCR Master Mix | Influenza B RT-PCR Master Mix |
|---|---|---|
| Nuclease free water | 280 µl | 280 µl |
| Influenza A primer mix | 28 µl | - |
| Influenza B primer mix | - | 28 µl |
| 2X Reaction Mix | 350 µl | 350 µl |
| SuperScript™ III RT/Platinum™ Taq Mix | 28 µl | 28 µl |
| Total volume | 686 µl | 686 µl |
For X24 samples, use 1.5 ml Eppendorf DNA LoBind tubes:
| Component | Influenza A RT-PCR Master Mix | Influenza B RT-PCR Master Mix |
|---|---|---|
| Nuclease free water | 560 µl | 560 µl |
| Influenza A primer mix | 56 µl | - |
| Influenza B primer mix | - | 56 µl |
| 2X Reaction Mix | 700 µl | 700 µl |
| SuperScript™ III RT/Platinum™ Taq Mix | 56 µl | 56 µl |
| Total volume | 1372 µl | 1372 µl |
For X48 samples, use 5 ml Eppendorf DNA LoBind tubes:
| Component | Influenza A RT-PCR Master Mix | Influenza B RT-PCR Master Mix |
|---|---|---|
| Nuclease free water | 1120 µl | 1120 µl |
| Influenza A primer mix | 112 µl | - |
| Influenza B primer mix | - | 112 µl |
| 2X Reaction Mix | 1400 µl | 1400 µl |
| SuperScript™ III RT/Platinum™ Taq Mix | 112 µl | 112 µl |
| Total volume | 2744 µl | 2744 µl |
For X96 samples, use 15 ml Eppendorf DNA LoBind tubes:
| Component | Influenza A RT-PCR Master Mix | Influenza B RT-PCR Master Mix |
|---|---|---|
| Nuclease free water | 2240 µl | 2240 µl |
| Influenza A primer mix | 224 µl | - |
| Influenza B primer mix | - | 224 µl |
| 2X Reaction Mix | 2800 µl | 2800 µl |
| SuperScript™ III RT/Platinum™ Taq Mix | 224 µl | 224 µl |
| Total volume | 5488 µl | 5488 µl |
For each influenza type, place a clean 96-well RT-PCR plate into a PCR-cooler or ice bucket with ice (if using).
Note: Enusre the RT-PCR plates for each influenza type are separate:
- One plate for Influenza A (Influenza A RT-PCR plate)
- One plate for Influenza B (Influenza B RT-PCR plate)
Using a stepper pipette or a multichannel pipette, aliquot 49 µl of influenza A RT-PCR Master Mix into the influenza A RT-PCR plate.
Using a stepper pipette or a multichannel pipette, aliquot 49 µl of influenza B RT-PCR Master Mix into the influenza B RT-PCR plate.
We recommend having a negative control for every plate of samples to monitor for contamination events.
Use 1 µl of nuclease-free water as your negative control input into a single well of each Influenza RT-PCR plate.
Seal the RT-PCR plate(s) and transfer to a template-addition pre-PCR hood.
Transfer 1 µl of influenza A samples to the wells containing influenza A RT-PCR Master Mix in the influenza A RT-PCR plate and mix thoroughly by pipetting the contents of each well up and down.
Transfer 1 µl of influenza B samples to the wells containing influenza B RT-PCR Master Mix in the influenza B RT-PCR plate and mix thoroughly by pipetting the contents of each well up and down.
Seal the RT-PCR plate(s) and spin down in a centrifuge.
Please note the thermal cycler programs are different for the Influenza A RT-PCR and Influenza B RT-PCR reactions.
Ensure you are using the correct program for your reaction plate.
Incubate the influenza A RT-PCR plate using the following program, with the heated lid set to 105°C:
| Step | Temperature | Time | Cycles |
|---|---|---|---|
| cDNA synthesis | 42°C | 60 min | 1 |
| Initial denaturation | 94°C | 2 min | 1 |
| Denaturation Annealing and extension | 94°C 45°C 68°C | 30 sec 30 sec 3 min | 5 |
| Denaturation Annealing and extension | 94°C 57°C 68°C | 30 sec 30 sec 3 min | 31 |
| Hold | 4°C | ∞ |
Incubate the influenza B RT-PCR plate using the following program, with the heated lid set to 105°C:
| Step | Temperature | Time | Cycles |
|---|---|---|---|
| cDNA synthesis | 45°C | 60 min | 1 |
| cDNA synthesis | 55°C | 30 min | 1 |
| Initial denaturation | 94°C | 2 min | 1 |
| Denaturation Annealing and extension | 94°C 40°C 68°C | 20 sec 30 sec 3 min 30 sec | 5 |
| Denaturation Annealing and extension | 94°C 58°C 68°C | 20 sec 30 sec 3 min 30 sec | 30 |
| Final extension | 68°C | 10 min | 1 |
| Hold | 4°C | ∞ |
If necessary, the protocol can be paused at this point. The samples should be kept at 4°C and can be stored overnight.
From this point onwards, a clean post-PCR hood can be used if available. Decontamination with UV and or DNAZap between sample batches is recommended.
Resuspend the AMPure XP beads by vortexing.
Add 50 µl of resuspended AMPure XP beads to each well of the RT-PCR plate(s) and mix by gently pipetting.
Incubate the RT-PCR plate(s) at room temperature for 10 minutes.
Prepare at least 500 µl 80% ethanol in nuclease-free water per sample.
Spin down the RT-PCR plate(s) and pellet the beads on a magnet for 5 minutes. Keep the plate on the magnet until the eluate is clear and colourless, and pipette off the supernatant.
Keep the plate on the magnet and wash the beads in each well with 200 µl of freshly prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Repeat the previous step.
Spin down and place the plate back on the magnet. Pipette off any residual ethanol. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the plate from the magnetic rack and resuspend each pellet in 15 µl nuclease-free water. Incubate for 2 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove and retain 15 µl of eluate containing the DNA per well, into a clean 96-well plate(s).
Dispose of the pelleted beads.
Quantify 1 µl of each eluted sample using a Qubit fluorometer.
Take forward your quantified samples to the end-prep step.
However, at this point it is also possible to store the samples at 4°C overnight.
For long-term storage or to store any unused amplified material for use in later experiments, store your samples at -20°C.
4. End-prep
耗材
- 无核酸酶水(如ThermoFisher,AM9937)
- NEBNext® Ultra™ II End Repair/dA-Tailing Module (E7546)
- 1.5 ml Eppendorf DNA LoBind 离心管
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Cat # 0030129504) with heat seals
仪器
- Multichannel pipette capable of dispensing 0.5 – 10 µL, and tips
- P1000 移液枪和枪头
- P200 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P10 移液枪和枪头
- 热循环仪
- 迷你离心机
- 盛有冰的冰桶
- 微孔板离心机
- 涡旋混匀仪
We recommended carrying the negative control through this step until sequencing.
Prepare the NEBNext Ultra II End Repair / dA-tailing Module reagents in accordance with manufacturer's instructions, and place on ice:
For optimal performance, NEB recommend the following:
Thaw all reagents on ice.
Ensure the reagents are well mixed.
Note: Do not vortex the Ultra II End Prep Enzyme Mix.
Always spin down tubes before opening for the first time each day.
The NEBNext Ultra II End Prep Reaction Buffer may contain a white precipitate. If this occurs, allow the mixture to come to room temperature and pipette the buffer several times to break up the precipitate, followed by a quick vortex to mix.
Determine the volume of the cleaned-up PCR reaction that yields 200 fmol of DNA per sample and aliquot into a clean 96-well plate (End-prep plate).
Make up each sample per well to 12.5 µl using nuclease-free water.
Prepare the following end-prep master mix in 1.5 ml Eppendorf DNA LoBind tube and mix thoroughly by pipetting:
| Reagent | Volume per reaction | For X24 samples | For X48 samples | For X96 samples |
|---|---|---|---|---|
| Ultra II End-prep Reaction Buffer | 1.75 µl | 52.5 µl | 105 µl | 210 µl |
| Ultra II End-prep Enzyme Mix | 0.75 µl | 22.5 µl | 45 µl | 90 µl |
| Total | 2.5 µl | 75 µl | 150 µl | 300 µl |
Using a stepper pipette or multi-channel pipette, add 2.5 µl of the end-prep master mix to each well containing 12.5 µl sample.
Ensure the reactions are thoroughly mixed by pipetting. Seal the End-prep plate and spin down briefly.
Using a thermal cycler, incubate the plate at 20°C for 5 minutes and 65°C for 5 minutes.
Take forward the end-prepped DNA into the native barcode ligation step.
If users want to pause the library preparation here, we recommend cleaning up your sample with 1X Agencourt AMPure XP beads and eluting in nuclease-free water before storing at 4°C.
5. Native barcode ligation
材料
- Native Barcodes (NB01-24) OR Native Barcodes (NB01-NB96)
- AMPure XP 磁珠(AXP)
- EDTA(EDTA)
- 短片段缓冲液(SFB)
耗材
- 新制备的 80% 乙醇(用无核酸酶水配制)
- 1.5 ml Eppendorf DNA LoBind 离心管
- 无核酸酶水(如 Thermo Scientific,AM9937)
- NEB Blunt/TA 连接酶预混液(NEB,M0367)
仪器
- Magnetic rack suitable for 96-well plates
- 热循环仪
- Hula混匀仪(低速旋转式混匀仪)
- 涡旋混匀仪
- 盛有冰的冰桶
- 迷你离心机
- P1000 移液枪和枪头
- P100 移液枪和枪头
- P10 移液枪和枪头
可选仪器
- Qubit™ 荧光计(或用于质控检测的等效仪器)
To monitor cross-contamination events, we recommend that the negative control is carried through this process and a barcode is used to sequence this control.
Prepare the NEB Blunt/TA Ligase Master Mix according to the manufacturer's instructions, and place on ice:
Thaw the reagents at room temperature.
Spin down the reagent tubes for 5 seconds.
Ensure the reagents are fully mixed by performing 10 full volume pipette mixes.
Thaw kit components at room temperature, then spin down briefly using a microfuge and mix as indicated in the Table below. Then place on ice:
| Reagent | 1. Thaw at room temperature | 2. Briefly spin down | 3. Mix by pipetting or vortexing | 4. Place on ice |
|---|---|---|---|---|
| EDTA (EDTA) | ✓ | ✓ | Vortexing | ✓ |
| Native Barcodes (NB01-24) or (NB01-96) | ✓ | ✓ | Only pipette mix immediately before use | ✓ |
| Short Fragment Buffer (SFB) | ✓ | ✓ | Vortexing | ✓ |
Select a unique barcode for each sample to be run together on the same flow cell.
Please note: Only use one barcode per sample.
条形码板孔仅限一次使用。使用前请确认所选孔密封完好;一旦刺穿或开启,不得再次使用。
Using a stepper pipette, or a multichannel pipette, make up the Native Barcode Ligation Plate in a clean 96-well plate. Add the reagents in the following order per well:
| Reagent | Volume per well for up to x24 samples | Volume per well for x25—x96 samples |
|---|---|---|
| Nuclease-free water | 6 µl | 3 µl |
| End-prepped DNA Note: Transfer to the corresponding well | 1.5 µl | 0.75 µl |
| Native barcode Note: Transfer to the corresponding well | 2.5 µl | 1.25 µl |
| Blunt/TA Ligation Master Mix | 10 µl | 5 µl |
| Total | 20 µl | 10 µl |
Depending on the number of samples, aliquot to each well of the columns:
| Plate location | x24 samples | x48 samples | x96 samples |
|---|---|---|---|
| Columns | 1-3 | 1-6 | 1-12 |
Mix the contents thoroughly by pipetting.
Seal the plate(s) and spin down briefly.
Incubate for 20 minutes at room temperature.
Add EDTA to each well and mix thoroughly by pipetting and spin down briefly.
| Up to x24 samples | x25 — x96 samples | |
|---|---|---|
| Volume of EDTA (blue cap) per sample | 4 µl | 2 µl |
EDTA is added at this step to stop the reaction.
Pool the barcoded samples in a clean 1.5 ml Eppendorf DNA LoBind tube:
| x24 samples | x48 samples | x96 samples | |
|---|---|---|---|
| Total volume for preps using EDTA (blue cap) | ~576 µl | ~576 µl | ~1,152 µl |
We recommend checking the base of your tubes/plate are all the same volume before pooling and after to ensure all the liquid has been taken forward.
Resuspend the AMPure XP Beads (AXP) by vortexing.
Add AMPure XP Beads (AXP) to the pooled reaction, and mix by pipetting for a 0.4X clean.
| Volume for 10 µl of sample | For 265 µl of samples | For 576 µl of samples | For 1,152 µl of samples | |
|---|---|---|---|---|
| Volume of AXP | 4 µl | 106 µl | 230 µl | 460 µl |
Incubate on a Hula mixer (rotator mixer) for 10 minutes at room temperature.
Prepare 500 µl of fresh 80% ethanol in nuclease-free water.
Spin down the sample and pellet the beads on a magnet for 5 minutes. Keep the tube on the magnet until the eluate is clear and colourless, and pipette off the supernatant.
Wash the beads by adding 700 μl Short Fragment Buffer (SFB). Flick the beads to resuspend, then return the tube to the magnetic rack and allow the beads to pellet. Remove the supernatant using a pipette and discard.
Repeat the previous step.
Keep the tube on the magnet and wash the beads with 100 µl of freshly prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Spin down and place the tube back on the magnet. Pipette off any residual ethanol. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend the pellet in 35 µl nuclease-free water by gently flicking.
Incubate for 10 minutes at 37°C. Every 2 minutes, agitate the sample by gently flicking for 10 seconds to encourage DNA elution.
Note: If stuggling to obtain the necessary yield, increasing the incubation period up to 30 minutes may improve elution efficacy.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove and retain 35 µl of eluate into a clean 1.5 ml Eppendorf DNA LoBind tube.
Quantify 1 µl of eluted sample using a Qubit fluorometer.
Take forward the barcoded DNA library to the adapter ligation and clean-up step. However, at this point it is also possible to store the sample at 4°C overnight.
6. Adapter ligation and clean-up
材料
- 短片段缓冲液(SFB)
- 洗脱缓冲液(EB)
- 免扩增接头(NA)
- AMPure XP 磁珠(AXP)
耗材
- NEBNext®快速连接模块(NEB,E6056)
- 1.5 ml Eppendorf DNA LoBind 离心管
- Qubit™ 分析管(Invitrogen, Q32856)
- Qubit™ dsDNA HS Assay(双链DNA高灵敏度检测)试剂盒(ThermoFisher,Q32851)
仪器
- 迷你离心机
- 磁力架
- 涡旋混匀仪
- Hula混匀仪(低速旋转式混匀仪)
- 热循环仪
- P1000 移液枪和枪头
- P200 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P10 移液枪和枪头
- 盛有冰的冰桶
- Qubit™ 荧光计(或用于质控检测的等效仪器)
The Native Adapter (NA) used in this kit and protocol is not interchangeable with other sequencing adapters.
测序芯片质检
我们强烈建议在开始文库制备之前,对测序芯片的活性纳米孔数量进行质检,以确保其足够支持实验的顺利进行。
详情请参阅 MinKNOW 实验指南中的 测序芯片质检说明。
Prepare the NEBNext Quick Ligation Reaction Module according to the manufacturer's instructions, and place on ice:
Thaw the reagents at room temperature.
Spin down the reagent tubes for 5 seconds.
Ensure the reagents are fully mixed by performing 10 full volume pipette mixes. Note: Do NOT vortex the Quick T4 DNA Ligase.
The NEBNext Quick Ligation Reaction Buffer (5x) may have a little precipitate. Allow the mixture to come to room temperature and pipette the buffer up and down several times to break up the precipitate, followed by vortexing the tube for several seconds to ensure the reagent is thoroughly mixed.
Do not vortex the Quick T4 DNA Ligase.
Spin down the Native Adapter (NA) and Quick T4 DNA Ligase, pipette mix and place on ice.
Thaw the Elution Buffer (EB) at room temperature and mix by vortexing. Then spin down and place on ice.
Thaw the Short Fragment Buffer (SFB) at room temperature and mix by vortexing. Then spin down and place on ice.
In a 1.5 ml Eppendorf LoBind tube, mix in the following order:
Between each addition, pipette mix 10 - 20 times.
| Reagent | Volume |
|---|---|
| Pooled barcoded sample | 30 µl |
| Native Adapter (NA) | 5 µl |
| NEBNext Quick Ligation Reaction Buffer (5X) | 10 µl |
| Quick T4 DNA Ligase | 5 µl |
| Total | 50 µl |
Thoroughly mix the reaction by gently pipetting and briefly spinning down.
Incubate the reaction for 20 minutes at room temperature.
The next clean-up step uses Short Fragment Buffer (SFB) and not 80% ethanol to wash the beads. The use of ethanol will be detrimental to the sequencing reaction.
Resuspend the AMPure XP Beads (AXP) by vortexing.
Add 20 µl of resuspended AMPure XP Beads (AXP) to the reaction and mix by pipetting.
Incubate on a Hula mixer (rotator mixer) for 10 minutes at room temperature.
Spin down the sample and pellet the beads on a magnet for 5 minutes. Keep the tube on the magnet until the eluate is clear and colourless, and pipette off the supernatant.
Wash the beads by adding 125 μl Short Fragment Buffer (SFB). Flick the beads to resuspend, then return the tube to the magnetic rack and allow the beads to pellet. Keep the tube on the magnet until the eluate is clear and colourless. Remove the supernatant using a pipette and discard.
Repeat the previous step.
Spin down and place the tube back on the magnet. Pipette off any residual supernatant. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend pellet in 15 µl Elution Buffer (EB).
Spin down and incubate for 10 minutes at 37°C. Every 2 minutes, agitate the sample by gently flicking for 10 seconds to encourage DNA elution.
Pellet the beads on a magnet until the eluate is clear and colourless, for at least 1 minute.
Remove and retain 15 µl of eluate containing the DNA library into a clean 1.5 ml Eppendorf DNA LoBind tube.
Dispose of the pelleted beads
Quantify 1 µl of eluted sample using a Qubit fluorometer.
Depending on your DNA library fragment size, prepare your final library in 12 µl of Elution Buffer (EB).
| Fragment library length | Flow cell loading amount |
|---|---|
| Very short (<1 kb) | 100 fmol |
| Short (1-10 kb) | 35–50 fmol |
| Long (>10 kb) | 300 ng |
Note: If the library yields are below the input recommendations, load the entire library.
If required, we recommend using a mass to mol calculator such as the NEB calculator.
The prepared library is used for loading onto the flow cell. Store the library on ice or at 4°C until ready to load.
Library storage recommendations
We recommend storing libraries in Eppendorf DNA LoBind tubes at 4°C for short-term storage or repeated use, for example, re-loading flow cells between washes. For single use and long-term storage of more than 3 months, we recommend storing libraries at -80°C in Eppendorf DNA LoBind tubes.
If quantities allow, the library may be diluted in Elution Buffer (EB) for splitting across multiple flow cells.
Depending on how many flow cells the library will be split across, more Elution Buffer (EB) than what is supplied in the kit will be required.
7. Priming and loading the MinION and GridION Flow Cell
材料
- 测序芯片冲洗液(FCF)
- 测序芯片系绳(FCT)
- 文库溶液(LIS)
- 文库颗粒(LIB)
- 测序缓冲液(SB)
耗材
- 1.5 ml Eppendorf DNA LoBind 离心管
- 无核酸酶水(如 Thermo Scientific,AM9937)
- 牛血清白蛋白(BSA)(50 mg/mL)(例如 Invitrogen™ UltraPure™ BSA 50 mg/mL, AM2616)
仪器
- MinION 或 GridION 测序仪
- SpotON Flow Cell
- MinION/GridION 测序芯片遮光片
- P1000 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P10 移液枪和枪头
Please note, this kit is only compatible with R10.4.1 flow cells (FLO-MIN114).
从冰箱中取出测序芯片,在室温下放置 20 分钟,以便在预处理和上样时更清晰地观察到传感器阵列。
Priming and loading a flow cell
We recommend all new users watch the 'Priming and loading your flow cell' video before your first run.
Using the Library Solution
For most sequencing experiments, use the Library Beads (LIB) for loading your library onto the flow cell. However, for viscous libraries it may be difficult to load with the beads and may be appropriate to load using the Library Solution (LIS).
Thaw the Sequencing Buffer (SB), Library Beads (LIB) or Library Solution (LIS, if using), Flow Cell Tether (FCT) and Flow Cell Flush (FCF) at room temperature before mixing by vortexing. Then spin down and store on ice.
For optimal sequencing performance and improved output on MinION R10.4.1 Flow Cells (FLO-MIN114), add Bovine Serum Albumin (BSA) to the flow cell priming mix at a final concentration of 0.2 mg/ml.
Note: We do not recommend using any other albumin type (e.g. recombinant human serum albumin).
To prepare the flow cell priming mix with BSA, combine Flow Cell Flush (FCF) and Flow Cell Tether (FCT), as directed below. Mix by pipetting at room temperature.
In a suitable tube for the number of flow cells, combine the following reagents:
| Reagent | Volume per flow cell |
|---|---|
| Flow Cell Flush (FCF) | 1,170 µl |
| Bovine Serum Albumin (BSA) at 50 mg/ml | 5 µl |
| Flow Cell Tether (FCT) | 30 µl |
| Total volume | 1,205 µl |
Open the MinION or GridION device lid and slide the flow cell under the clip. Press down firmly on the priming port cover to ensure correct thermal and electrical contact.
Complete a flow cell check to assess the number of pores available before loading the library.
This step can be omitted if the flow cell has been checked previously.
See the flow cell check document for more information.
Slide the flow cell priming port cover clockwise to open the priming port.
Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µl, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the priming port
- Turn the wheel until the dial shows 220-230 µl, to draw back 20-30 µl, or until you can see a small volume of buffer entering the pipette tip
Note: Visually check that there is continuous buffer from the priming port across the sensor array.
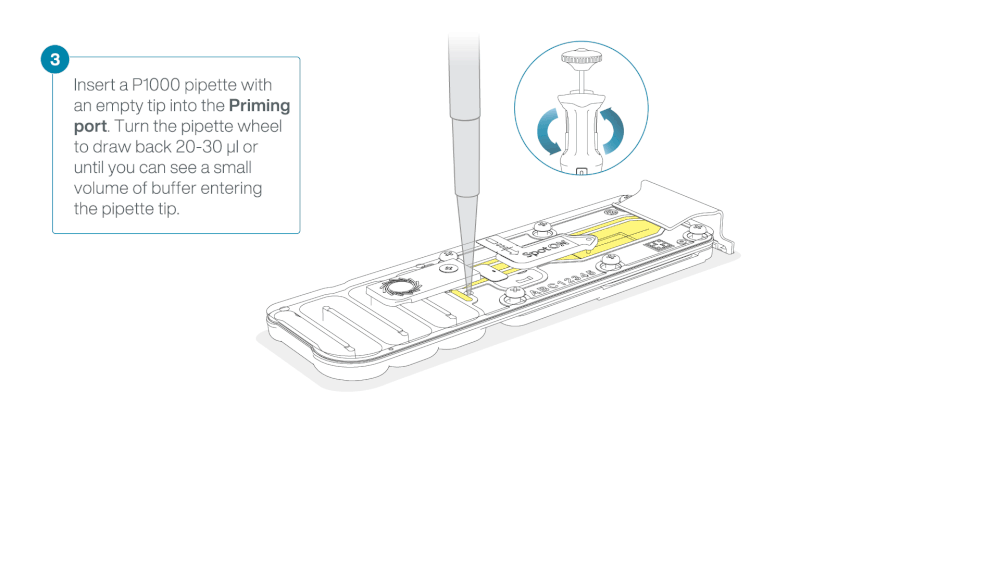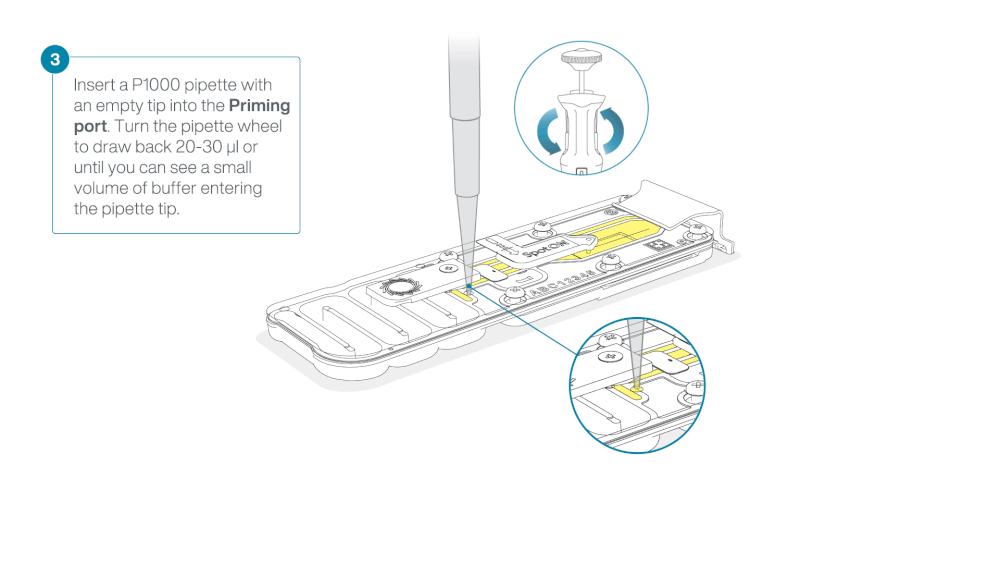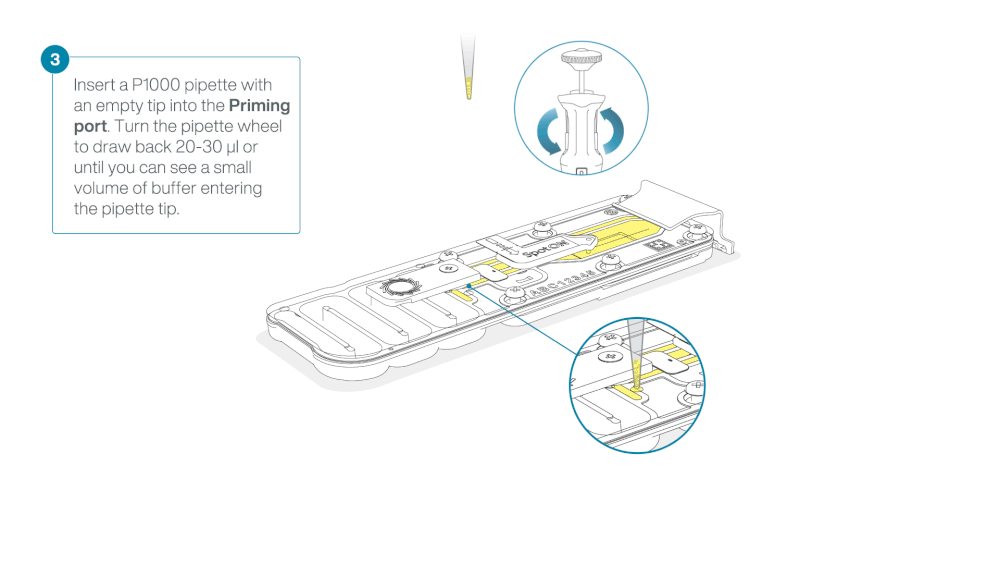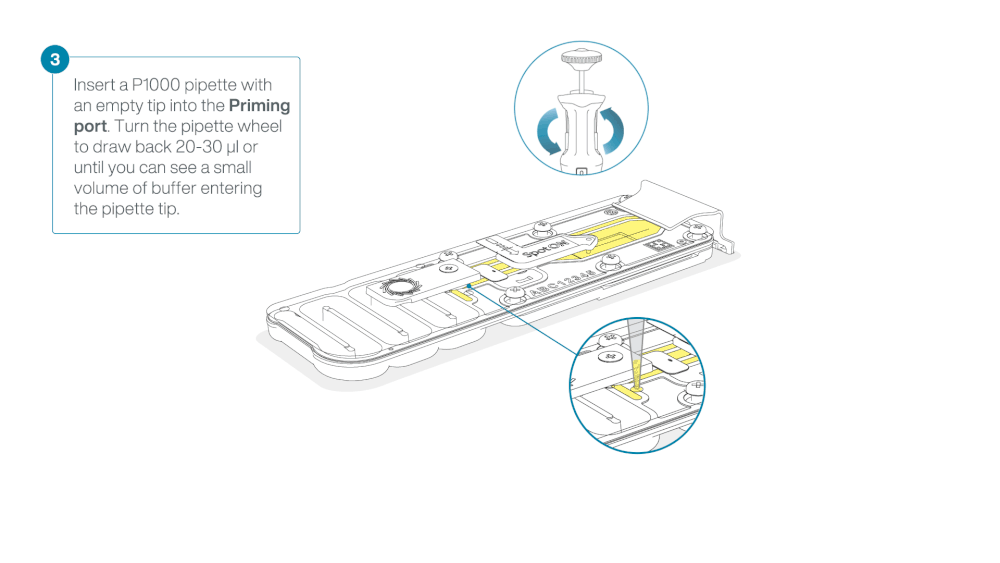
Load 800 µl of the priming mix into the flow cell via the priming port, avoiding the introduction of air bubbles. Wait for five minutes. During this time, prepare the library for loading by following the steps below.
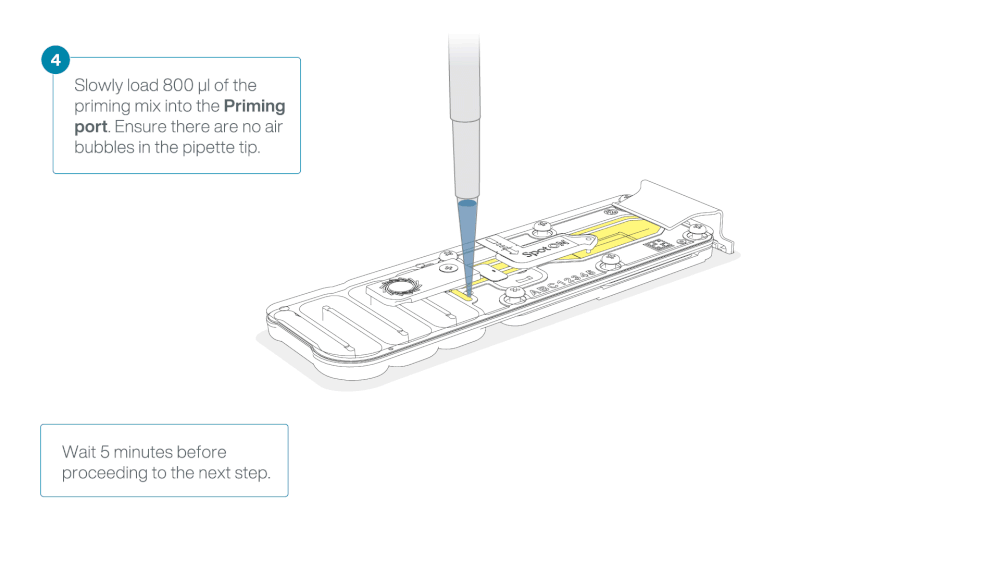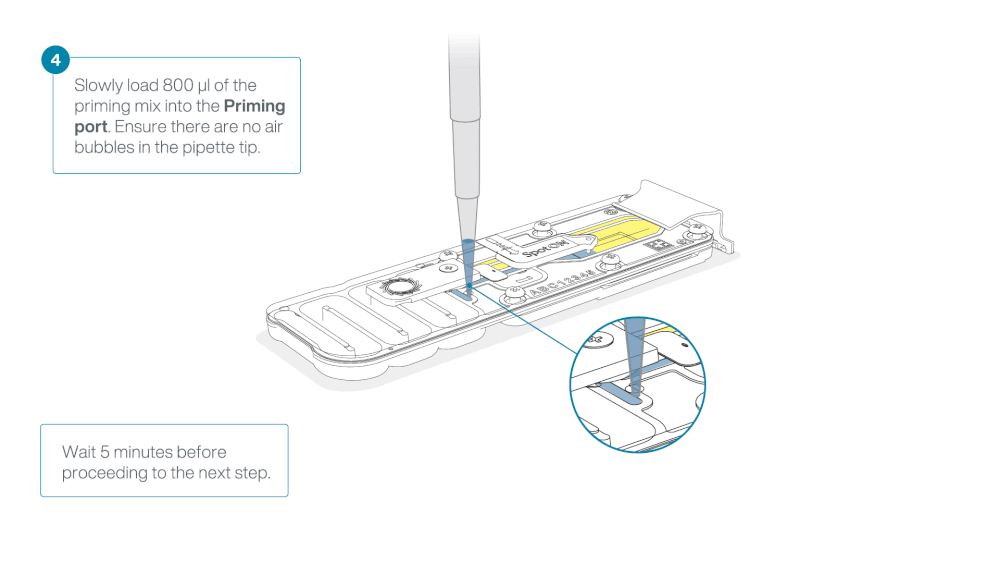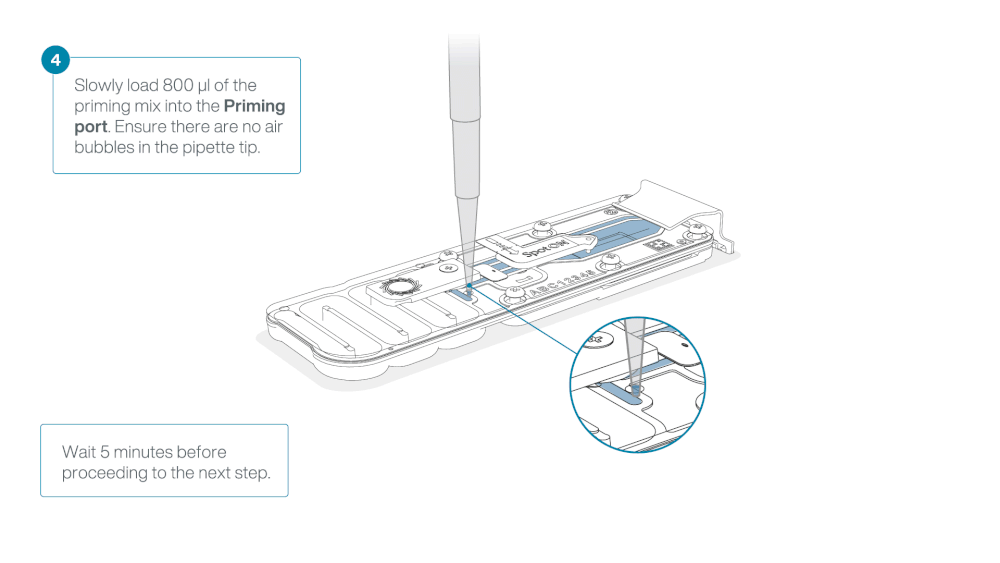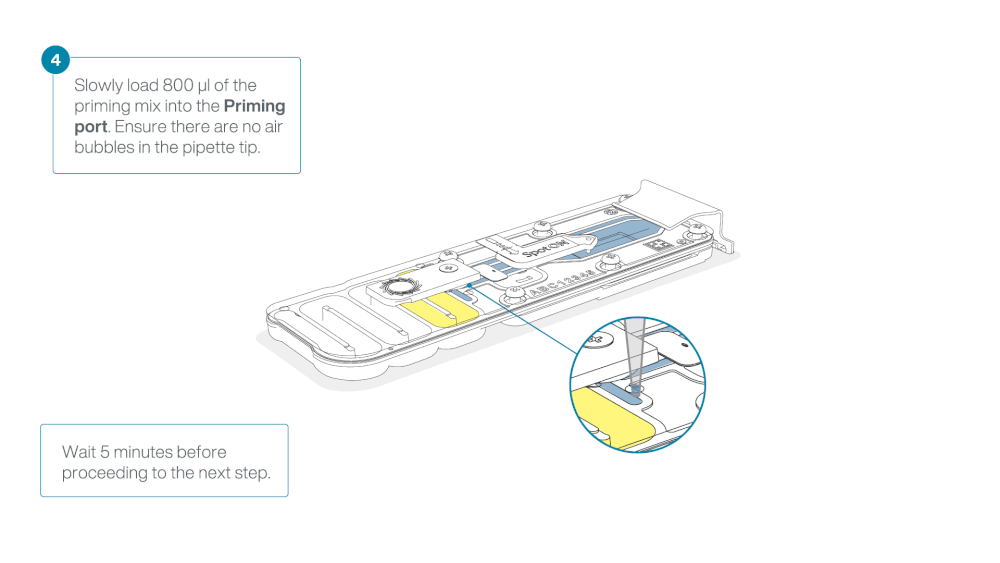
Thoroughly mix the contents of the Library Beads (LIB) by pipetting.
The Library Beads (LIB) tube contains a suspension of beads. These beads settle very quickly. It is vital that they are mixed immediately before use.
We recommend using the Library Beads (LIB) for most sequencing experiments. However, the Library Solution (LIS) is available for more viscous libraries.
In a new 1.5 ml Eppendorf DNA LoBind tube, prepare the library for loading as follows:
| Reagent | Volume per flow cell |
|---|---|
| Sequencing Buffer (SB) | 37.5 µl |
| Library Beads (LIB) mixed immediately before use, or Library Solution (LIS), if using | 25.5 µl |
| DNA library | 12 µl |
| Total | 75 µl |
Complete the flow cell priming:
- Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
- Load 200 µl of the priming mix into the flow cell priming port (not the SpotON sample port), avoiding the introduction of air bubbles.

Mix the prepared library gently by pipetting up and down just prior to loading.
Add 75 μl of the prepared library to the flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.

Gently replace the SpotON sample port cover, making sure the bung enters the SpotON port and close the priming port.

For optimal sequencing output, install the light shield on your flow cell as soon as the library has been loaded.
We recommend leaving the light shield on the flow cell when library is loaded, including during any washing and reloading steps. The shield can be removed when the library has been removed from the flow cell.
Place the light shield onto the flow cell, as follows:
Carefully place the leading edge of the light shield against the clip. Note: Do not force the light shield underneath the clip.
Gently lower the light shield onto the flow cell. The light shield should sit around the SpotON cover, covering the entire top section of the flow cell.

The MinION Flow Cell Light Shield is not secured to the flow cell and careful handling is required after installation.
Close the device lid and set up a sequencing run on MinKNOW.
When a flow cell is inserted into the MinION Mk1D, the device lid will sit on top of the flow cell, leaving a small gap around the sides. This is normal and has no impact on the performance of the device.
Please refer to this FAQ regarding the device lid.

8. Data acquisition and basecalling
We do not recommend sequencing and performing data analysis simultaneously on your device.
To ensure the compute on your device can keep up with the requirements for sequencing and/or analysis, we strongly recommend against running both processes at the same time.
Ensure your sequencing run has completed before setting off data analysis. Data analysis will be performed post-sequencing.
Equally, we do not recommend starting a sequencing run if you are currently performing data analysis on your device.
How to start sequencing
The sequencing device control, data acquisition and real-time basecalling are carried out by the MinKNOW software. Please ensure MinKNOW is installed on your computer or device. Further instructions for setting up a sequencing run can be found in the MinKNOW protocol.
We recommend setting up your sequencing run using the basecalling and barcoding recommendations outlined below. All other parameters can be left to their default settings.
MinKNOW settings
For fastest turnaround time and easiest analysis, basecalling should be performed live during the sequencing run using the High Accuracy (HAC) basecaller.
Below are the recommendations for MinKNOW settings:
Positions
Flow cell position: [user defined]
Experiment name: [user defined]
Flow cell type: FLO-MIN114
Sample ID: [user defined]
Kit
Kit selection: Native Barcoding Sequencing Kit 24 (SQK-NBD114.24) or Native Barcoding Sequencing Kit 96 (SQK-NBD114.96)
Run configuration
Sequencing and analysis
Basecalling: On [default]
Modified bases: Off
Model: High-accuracy basecalling (HAC) [default]
Barcoding: On [default]
Trim barcodes: Off [default]
Barcode both ends: On
Custom barcodes selection: Off [default]
Alignment: Off [default]
Adaptive sampling: Off [default]
Advanced options
Active channel selection: On [default]
Time between pore scans: 1.5 [default]
Reserve pores: On [default]
Data targets
Run limit: [user defined]*
*Sequencing time will depend on data requirements.
Output
Output format
.POD5: On [default]
.FASTQ: On [default]
.BAM: On
Filtering: On [default]
Qscore: 9 [default]
Minimum read length: 200 bp [default]
9. Flow cell reuse and returns
材料
- 测序芯片清洗剂盒(EXP-WSH004)
After your sequencing experiment is complete, if you would like to reuse the flow cell, please follow the Flow Cell Wash Kit protocol and store the washed flow cell at +2°C to +8°C.
The Flow Cell Wash Kit protocol is available on the Nanopore Community.
We recommend you to wash the flow cell as soon as possible after you stop the run. However, if this is not possible, leave the flow cell on the device and wash it the next day.
Alternatively, follow the returns procedure to send the flow cell back to Oxford Nanopore.
Instructions for returning flow cells can be found here.
If you encounter issues or have questions about your sequencing experiment, please refer to the Troubleshooting Guide in this protocol.
10. Downstream analysis
Recommended analysis pipeline
The analysis of the FASTQ format sequence data is performed using a Nextflow workflow called Influenza Typing Workflow (wf-flu). The use of the Nextflow software has been integrated into the EPI2ME Labs software that we recommend for running our downstream analysis methods.
Alternative methods for downstream analysis are available using your device terminal or command line, however we only suggest this for experienced users.
The workflow processes the basecalled and demultiplexed DNA sequence data output generated by MinKNOW:
- The sequences are filtered for a minimum length and quality thresholds (200 nucleotides and Q9 respectively) prior to sequence alignment to the CDC multi-fasta Influenza reference.
- The alignment is performed using the Minimap2 software.
- Depth of coverage across the mapped sequences is measured using Samtools before genetic variants are called using Medaka.
- A coverage-masked consensus sequence is prepared for each sample using bcftools.
- The influenza strain typing is then performed using the abricate software with an insaflu database.
The influenza strains included in the database are listed in the project documentation pages for the Influenza Typing Workflow.
The workflow returns a per-run HTML-format summary report along with a CSV file of typing results. Additional files that include mapping BAM files and VCF files of Medaka variants are also included in the workflow output.
For more information, please refer to the Influenza workflow blog.
Software set-up and installation
The EPI2ME application provides a clean interface to accessing bioinformatics workflows, and is our recommended method in performing your post-sequencing analysis.
Follow the instructions in the EPI2ME Installation guide to install the application on your device.
For more information on how to use EPI2ME, refer to the EPI2ME Quick Start guide.
Installing and updating the wf-flu workflow in EPI2ME Labs:
Ensure you have installed the wf-flu workflow prior to the first analysis set-up.
In the EPI2ME Labs home page, scroll down to the "Install workflows" section and click on epi2me-labs/wf-flu:

If you have already installed the wf-flu workflow, ensure you are using the latest version.
Updating the workflow can be done directly through EPI2ME Labs by navigating to the wf-flu workflow page and clicking Update Workflow:

Demultiplexing of multiple barcoded samples
The wf-flu analysis requires FASTQ sequence data that has already been demultiplexed.
Reads will be demultiplexed during sequencing if you are following the recommended "Required settings in MinKNOW". However, demultiplexing can also be done post-sequencing using the MinKNOW software.
For more information and guides on demultiplexing using MinKNOW, refer to the "Post-run analysis" section in our MinKNOW Protocol.
The expected input for wf-flu is a folder of folders as shown below. Each of the barcode folders should contain the FASTQ sequence data and files may either be uncompressed or gzipped.
$ tree -d FluFastq/
FluFastq/
├── barcode01
├── barcode02
├── barcode03
├── barcode04
├── barcode05
├── barcode06
└── unclassified
Basecalling model
The basecalling model should be specified when setting up the wf-flu analysis. This should reflect the basecalling model selected during your run set-up as follows:
- If using the default model, High-accuracy basecalling (HAC): r1041_e82_400bps_hac_variant_g615
- If you have used Super accurate basecalling (SUP), please use: r1041_e82_400bps_sup_variant_g615
- If you have used FAST basecalling, please use: r1041_e82_400bps_fast_variant_g615
Running a Flu analysis using EPI2ME Labs
Open the EPI2ME Labs application on your device.
Open the "Workflows" tab in the EPI2ME Labs application and click on the "wf-flu" workflow:

In the "wf-flu" workflow page, select "Run this workflow" to open analysis set-up:

Complete the wf-flu run set-up:
Select your data input file location. Please note, this folder must contain the demultiplexed FASTQ files of your sequencing run.
Set the basecaller cgf to the basecalling model used in your sequencing run.

Expand the Extra configuration tab and set the Run name for your wf-flu analysis.

Click Launch workflow at the bottom of the page to begin your analysis.
Completed analysis and result files
The wf-flu analysis outputs will be written to the Working Directory folder specified in the EPI2ME Labs Settings tab.
The location of this folder is specified in the wf-flu run Instance parameters preceeded by out_dir.
However, these files can also be accessed directly in the EPI2ME Labs application from the completed analysis page for your run:

Housekeeping and disk usage
The "Working Directory" can be specified in the EPI2ME Labs "Settings" tab and defines where the workflow intermediate files and outputs are stored.
This folder will accumulate a significant number of files that correspond to raw BAM files, other larger intermediates and analysis results files. We recommend this folder to be routinely cleared.
11. Issues during DNA/RNA extraction and library preparation for Kit 14
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Low sample quality
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) | The DNA extraction method does not provide the required purity | The effects of contaminants are shown in the Contaminants document. Please try an alternative extraction method that does not result in contaminant carryover. Consider performing an additional SPRI clean-up step. |
| Low RNA integrity (RNA integrity number <9.5 RIN, or the rRNA band is shown as a smear on the gel) | The RNA degraded during extraction | Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. |
| RNA has a shorter than expected fragment length | The RNA degraded during extraction | Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. We recommend working in an RNase-free environment, and to keep your lab equipment RNase-free when working with RNA. |
Low DNA recovery after AMPure bead clean-up
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low recovery | DNA loss due to a lower than intended AMPure beads-to-sample ratio | 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample. 2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up. |
| Low recovery | DNA fragments are shorter than expected | The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 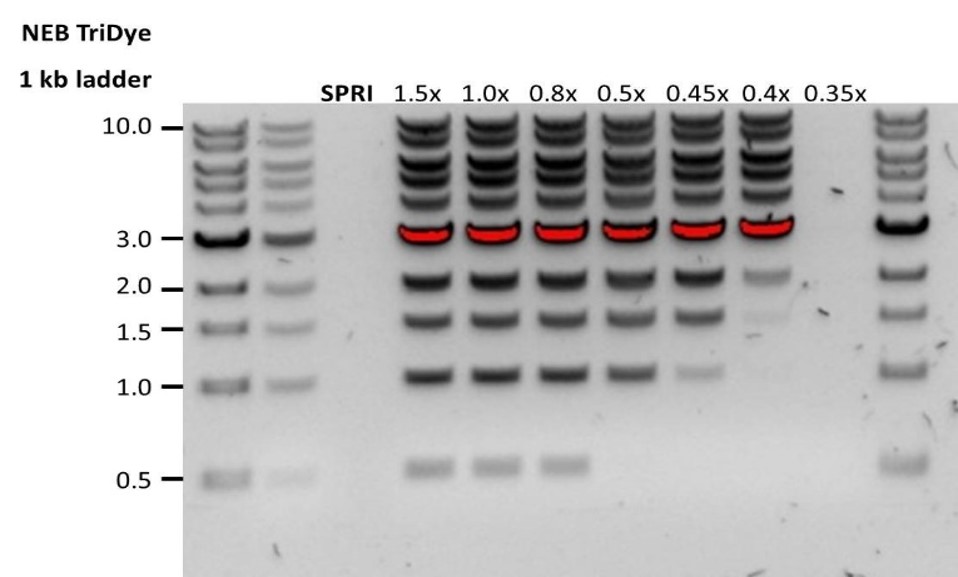 |
| Low recovery after end-prep | The wash step used ethanol <70% | DNA will be eluted from the beads when using ethanol <70%. Make sure to use the correct percentage. |
12. Issues during the sequencing run for Kit 14
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Fewer pores at the start of sequencing than after Flow Cell Check
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | An air bubble was introduced into the nanopore array | After the Flow Cell Check it is essential to remove any air bubbles near the priming port before priming the flow cell. If not removed, the air bubble can travel to the nanopore array and irreversibly damage the nanopores that have been exposed to air. The best practice to prevent this from happening is demonstrated in this video. |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | The flow cell is not correctly inserted into the device | Stop the sequencing run, remove the flow cell from the sequencing device and insert it again, checking that the flow cell is firmly seated in the device and that it has reached the target temperature. If applicable, try a different position on the device (GridION/PromethION). |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | Contaminations in the library damaged or blocked the pores | The pore count during the Flow Cell Check is performed using the QC DNA molecules present in the flow cell storage buffer. At the start of sequencing, the library itself is used to estimate the number of active pores. Because of this, variability of about 10% in the number of pores is expected. A significantly lower pore count reported at the start of sequencing can be due to contaminants in the library that have damaged the membranes or blocked the pores. Alternative DNA/RNA extraction or purification methods may be needed to improve the purity of the input material. The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
MinKNOW script failed
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Script failed" | Restart the computer and then restart MinKNOW. If the issue persists, please collect the MinKNOW log files and contact Technical Support. If you do not have another sequencing device available, we recommend storing the flow cell and the loaded library at 4°C and contact Technical Support for further storage guidance. |
Pore occupancy below 40%
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Pore occupancy <40% | Not enough library was loaded on the flow cell | 10–20 fmol of good quality library can be loaded on to a MinION/GridION flow cell. Please quantify the library before loading and calculate mols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to pmol" |
| Pore occupancy close to 0 | The Native Barcoding Kit was used, and ethanol was used instead of LFB or SFB at the wash step after sequencing adapter ligation | Ethanol can denature the motor protein on the sequencing adapters. Make sure the LFB or SFB buffer was used after ligation of sequencing adapters. |
| Pore occupancy close to 0 | No tether on the flow cell | Tethers are adding during flow cell priming (FCT tube). Make sure FCT was added to FCF before priming. |
Shorter than expected read length
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Shorter than expected read length | Unwanted fragmentation of DNA sample | Read length reflects input DNA fragment length. Input DNA can be fragmented during extraction and library prep. 1. Please review the Extraction Methods in the Nanopore Community for best practice for extraction. 2. Visualise the input DNA fragment length distribution on an agarose gel before proceeding to the library prep. 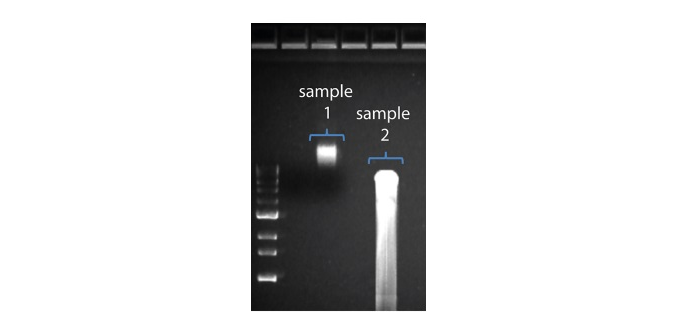 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented. In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.3. During library prep, avoid pipetting and vortexing when mixing reagents. Flicking or inverting the tube is sufficient. |
Large proportion of unavailable pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
Large proportion of unavailable pores (shown as blue in the channels panel and pore activity plot) 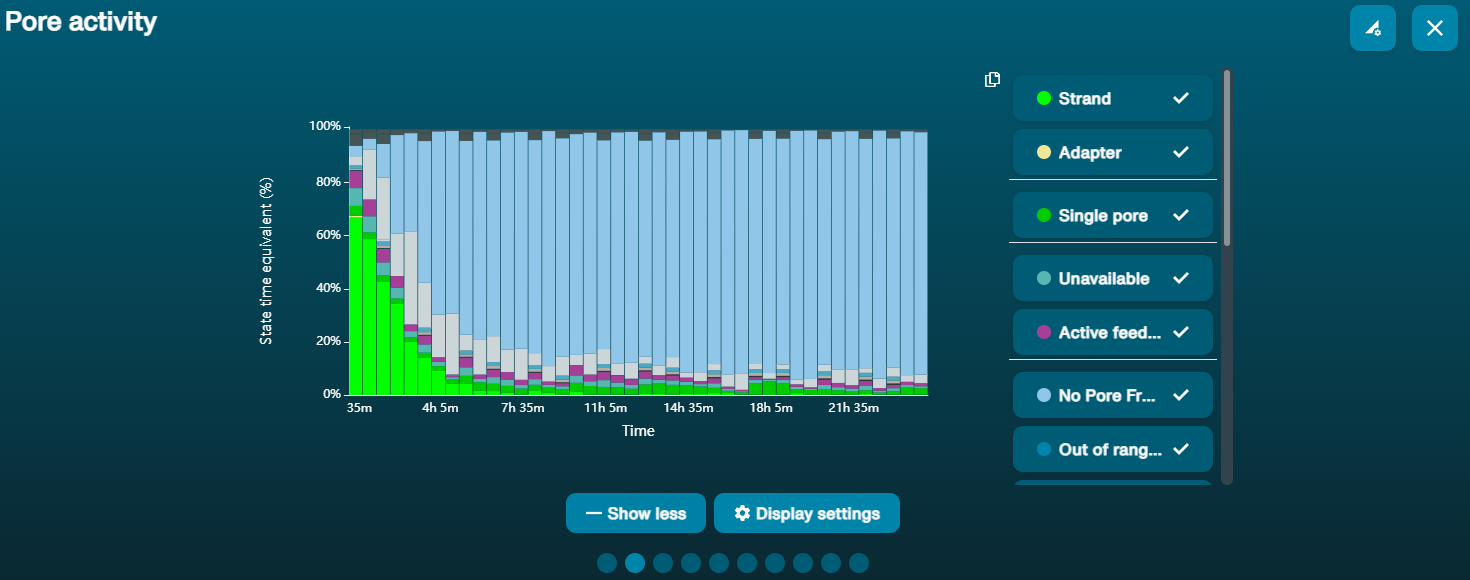 The pore activity plot above shows an increasing proportion of "unavailable" pores over time. The pore activity plot above shows an increasing proportion of "unavailable" pores over time. | Contaminants are present in the sample | Some contaminants can be cleared from the pores by the unblocking function built into MinKNOW. If this is successful, the pore status will change to "sequencing pore". If the portion of unavailable pores stays large or increases: 1. A nuclease flush using the Flow Cell Wash Kit (EXP-WSH004) can be performed, or 2. Run several cycles of PCR to try and dilute any contaminants that may be causing problems. |
Large proportion of inactive pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Large proportion of inactive/unavailable pores (shown as light blue in the channels panel and pore activity plot. Pores or membranes are irreversibly damaged) | Air bubbles have been introduced into the flow cell | Air bubbles introduced through flow cell priming and library loading can irreversibly damage the pores. Watch the Priming and loading your flow cell video for best practice |
| Large proportion of inactive/unavailable pores | Certain compounds co-purified with DNA | Known compounds include polysaccharides. 1. Clean-up using the QIAGEN PowerClean Pro kit. 2. Perform a whole genome amplification with the original gDNA sample using the QIAGEN REPLI-g kit. |
| Large proportion of inactive/unavailable pores | Contaminants are present in the sample | The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
Reduction in sequencing speed and q-score later into the run
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Reduction in sequencing speed and q-score later into the run | Fast fuel consumption is typically seen in Kit 9 chemistry (e.g. SQK-LSK109) when the flow cell is overloaded with library. Please see the appropriate protocol for your DNA library to find the recommendation. | Add more fuel to the flow cell by following the instructions in the MinKNOW protocol. In future experiments, load lower amounts of library to the flow cell. |
Temperature fluctuation
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Temperature fluctuation | The flow cell has lost contact with the device | Check that there is a heat pad covering the metal plate on the back of the flow cell. Re-insert the flow cell and press it down to make sure the connector pins are firmly in contact with the device. If the problem persists, please contact Technical Services. |
Failed to reach target temperature
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Failed to reach target temperature" | The instrument was placed in a location that is colder than normal room temperature, or a location with poor ventilation (which leads to the flow cells overheating) | MinKNOW has a default timeframe for the flow cell to reach the target temperature. Once the timeframe is exceeded, an error message will appear and the sequencing experiment will continue. However, sequencing at an incorrect temperature may lead to a decrease in throughput and lower q-scores. Please adjust the location of the sequencing device to ensure that it is placed at room temperature with good ventilation, then re-start the process in MinKNOW. Please refer to this link for more information on MinION temperature control. |







