Direct-from-colony rapid microbial sequencing for Salmonella serotyping using SQK-RPB114.24 (DCM_9210_V114_revG_28Apr2026)
MinION: Protocol
Direct-from-colony rapid microbial sequencing for Salmonella serotyping using SQK-RPB114.24 V DCM_9210_V114_revG_28Apr2026
This is a rapid method to perform sequencing and analysis of Salmonella.
The protocol:
- Uses Salmonella colonies from overnight cultures
- Includes tagmentation, barcoding, and PCR amplification
- Enables multiplexing of 1-24 samples
- Is compatible with R10.4.1 flow cells
For Research Use Only
FOR RESEARCH USE ONLY.
Contents
Introduction to the protocol
Library preparation
Sequencing and data analysis
Troubleshooting
Overview
This is a rapid method to perform sequencing and analysis of Salmonella.
The protocol:
- Uses Salmonella colonies from overnight cultures
- Includes tagmentation, barcoding, and PCR amplification
- Enables multiplexing of 1-24 samples
- Is compatible with R10.4.1 flow cells
For Research Use Only
1. Overview of the protocol
This is an Early Access product
For more information about our Early Access programmes, please see this article on product release phases.
Please ensure you always use the most recent version of the protocol.
Introduction to the direct-from-colony microbial sequencing: rapid Salmonella serotyping protocol
This protocol describes a method for preparing, sequencing, and analysis of up to 24 Salmonella colonies using the Rapid PCR Barcoding Kit V14 (SQK-RPB114.24).
This simple end-to-end solution allows you to generate whole genome sequencing data directly from a single Salmonella colony, eliminating the need for subculture into liquid broth, DNA extraction or complex library preparation methods, thereby offering faster turn-around times.
Combined with our custom wf-bacterial-genomes analysis pipeline using EPI2ME, this method provides an organism species ID, serotype, multi locus sequence type (MLST) and antibiotic resistance (AMR) profile information.
This method is designed to provide non-circular genomes for research and routine lab operations, including food safety testing purposes.
For more information and additional options for this end-to-end workflow, please visit our know-how document.

Steps in the end-to-end workflow:
Prepare for your experiment
You will need to:
Ensure pathogenic organisms are handled in appropriate biosafety conditions. Please adhere to the correct health and safety practices in accordance to your laboratory standards, and local rules and regulations.
This method should be performed in the appropriate containment level laboratory spaces (biosafety cabinets: BSC) until the sample tagmentation stage. Following sample tagmentation, cells are non-viable and can be used outside the biosafety cabinet.Streak your sample onto a freshly prepared agar plate appropriately such that single Salmonella colonies can be isolated following an overnight incubation. We recommend the use of TSA plates for culturing Salmonella.
The quality checks performed during the protocol are essential in ensuring experimental success.Ensure you have your sequencing kit, the correct equipment, and third-party reagents.
Download the software for acquiring and analysing your data.
Check your flow cell to ensure it has enough pores for a good sequencing run.
Library preparation
The Table below is an overview of the steps required in the library preparation, including timings and optional stopping points.
| Step | Process | Time | Stop option |
|---|---|---|---|
| Colony growth | Grow Salmonella single colony culture(s) of ~2-3 mm overnight on an agar plate. | ~16-18 hours | |
| Sample preparation | Resuspend each single colony in Tris buffer. | 5 minutes | |
| DNA tagmentation | Tagment your DNA using the Fragmentation Mix from the sequencing kit. | 10 minutes | |
| PCR amplification | PCR your sample using the barcoded primer supplied in the sequencing kit. | 180 minutes | We do not recommend leaving the PCR reaction overnight. Using the amplified samples straight after PCR will yield better results. If sample storage is unavoidable, add 4 µl EDTA to each sample and then store at 4°C. |
| Sample pooling and clean-up | Pool your barcoded PCR samples in equal ratios and perform a clean-up. | 30 minutes | 4°C overnight |
| Rapid adapter attachment and clean-up | Attach the sequencing adapters to the DNA ends. | 5 minutes | We strongly recommend sequencing your library as soon as it is adapted. |
| Priming and loading the flow cell | Prime the flow cell and load the prepared library for sequencing. | 10 minutes |
Sequencing and analysis
You will need to:
Start a sequencing run using the MinKNOW software, which will collect raw data from the device and convert it into basecalled reads.
Start the EPI2ME software and select the wf-bacterial-genomes workflow.
Compatibility of this protocol
This protocol should only be used in combination with:
- Rapid PCR Barcoding Kit 24 V14 (SQK-RPB114.24)
- R10.4.1 flow cells (FLO-MIN114)
- Flow Cell Wash Kit (EXP-WSH004)
- Rapid Adapter Auxiliary V14 (EXP-RAA114)
- Sequencing Auxiliary Vials V14 (EXP-AUX003)
- Flow Cell Priming Kit V14 (EXP-FLP004)
- MinION Mk1D: MinION Mk1D – device and IT specifications
- GridION: GridION Mk1 – device and IT specifications
2. Equipment and consumables
Materials
- Up to 24x single Salmonella colonies (2–3 mm in diameter)
- Rapid PCR Barcoding Kit 24 V14 (SQK-RPB114.24)
Consumables
- MinION/GridION Flow Cell
- 1 M Tris Buffer, pH8 (e.g. Invitrogen™, AM9855G)
- LongAmp Hot Start Taq 2X Master Mix (NEB, M0533)
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Bovine Serum Albumin (BSA) (50 mg/ml) (e.g. Invitrogen™ UltraPure™ BSA 50 mg/ml, AM2616)
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- 1.5 ml Eppendorf DNA LoBind tubes
- 0.2 ml thin-walled PCR tubes
Equipment
- MinION or GridION device
- MinION/GridION Flow Cell Light Shield
- Inoculation loop or equivalent for picking your colony
- Ice bucket with ice
- Microfuge
- Timer
- Thermal cycler
- Magnetic separation rack
- Hula mixer (gentle rotator mixer)
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P2 pipette and tips
- Multichannel pipette and tips
- Qubit™ fluorometer (or equivalent for QC check)
Optional equipment
- Agilent Bioanalyzer (or equivalent)
- Microbiological Safety Cabinet II contamination level 3
For this protocol, you will need a single colony of Salmonella 2–3 mm in diameter for each input.
Streak your sample onto a freshly prepared agar plate appropriately such that single Salmonella colonies can be isolated following an overnight incubation. We recommend the use of TSA plates for culturing Salmonella.
Optional: Alternatively, you can use 3-5 ng in 3 μl volume per sample of isolated DNA as input for library preparation.
Input DNA
How to QC your input DNA
It is important that the input DNA meets the quantity and quality requirements. Using too little or too much DNA, or DNA of poor quality (e.g. highly fragmented or containing RNA or chemical contaminants) can affect your library preparation.
For instructions on how to perform quality control of your DNA sample, please read the Input DNA/RNA QC protocol.
Chemical contaminants
Depending on how the DNA is extracted from the raw sample, certain chemical contaminants may remain in the purified DNA, which can affect library preparation efficiency and sequencing quality. Read more about contaminants on the Contaminants page of the Community.
Third-party reagents
We have validated and recommend the use of all the third-party reagents used in this protocol. Alternatives have not been tested by Oxford Nanopore Technologies.
For all third-party reagents, we recommend following the manufacturer's instructions to prepare the reagents for use.
Check your flow cell
We highly recommend that you check the number of pores in your flow cell prior to starting a sequencing experiment. This should be done within 12 weeks of purchasing your MinION/GridION Flow Cells. Oxford Nanopore Technologies will replace any unused flow cell with fewer than the number of pores listed in the Table below, when the result is reported within two days of performing the flow cell check, and when the storage recommendations have been followed. To do the flow cell check, please follow the instructions in the Flow Cell Check document.
| Flow cell | Minimum number of active pores covered by warranty |
|---|---|
| MinION/GridION Flow Cell | 800 |
The Rapid Adapter (RA) used in this kit and protocol is not interchangeable with other sequencing adapters.
Rapid PCR Barcoding Kit 24 V14 (SQK-RPB114.24) contents
We are in the process of reformatting the Rapid Barcode Primers provided in this kit into a plate format. Additionally we have lowered the concentration of the Rapid Barcode Primers in the plate format to 1 µM (down from 10 µM in the vial format). These changes will reduce plastic waste and facilitates automated applications.
Please ensure you follow the correct instructions in the method for your kit format and Rapid Barcode Primers concentration.
Plate format with Rapid Barcode Primers at 1 µM
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| Fragmentation Mix | FRM | 1 | Brown | 160 |
| Rapid Adapter | RA | 1 | Green | 15 |
| Adapter Buffer | ADB | 1 | Clear | 100 |
| AMPure XP Beads | AXP | 3 | Amber | 1,200 |
| Elution Buffer | EB | 2 | Black | 500 |
| EDTA | EDTA | 1 | Blue | 700 |
| Sequencing Buffer | SB | 1 | Red | 700 |
| Library Beads | LIB | 1 | Pink | 600 |
| Library Solution | LIS | 1 | White cap, pink label | 600 |
| Flow Cell Flush | FCF | 1 | Clear cap, light blue label | 8,000 |
| Flow Cell Tether | FCT | 1 | Purple | 200 |
| Rapid Barcode Primers (01-24) at 1 µM | RPB | - | 2 plates, 3 sets of primer barcodes per plate | 15 µl per well |
Note: This product contains AMPure XP reagent manufactured by Beckman Coulter, Inc. and can be stored at -20°C with the kit without detriment to reagent stability.
Vial format with Rapid Barcode Primers at 10 µM
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| Fragmentation Mix | FRM | Brown | 1 | 160 |
| Rapid Adapter | RA | Green | 1 | 15 |
| Adapter Buffer | ADB | Clear | 1 | 100 |
| AMPure XP Beads | AXP | Amber | 3 | 1,200 |
| Elution Buffer | EB | Black | 2 | 500 |
| EDTA | EDTA | Blue | 1 | 700 |
| Sequencing Buffer | SB | Red | 1 | 700 |
| Library Beads | LIB | Pink | 1 | 600 |
| Library Solution | LIS | White cap, pink label | 1 | 600 |
| Flow Cell Flush | FCF | Clear | 1 | 8,000 |
| Flow Cell Tether | FCT | Purple | 1 | 200 |
| Rapid Barcode Primer 01-24 | RLB01-24 | Clear | 24 (one per barcode) | 15 |
Note: This product contains AMPure XP reagent manufactured by Beckman Coulter, Inc. and can be stored at -20°C with the kit without detriment to reagent stability.
Rapid Barcode Primers
| Component | Sequence |
|---|---|
| Rapid Barcode Primer 01 | AAGAAAGTTGTCGGTGTCTTTGTG |
| Rapid Barcode Primer 02 | TCGATTCCGTTTGTAGTCGTCTGT |
| Rapid Barcode Primer 03 | GAGTCTTGTGTCCCAGTTACCAGG |
| Rapid Barcode Primer 04 | TTCGGATTCTATCGTGTTTCCCTA |
| Rapid Barcode Primer 05 | CTTGTCCAGGGTTTGTGTAACCTT |
| Rapid Barcode Primer 06 | TTCTCGCAAAGGCAGAAAGTAGTC |
| Rapid Barcode Primer 07 | GTGTTACCGTGGGAATGAATCCTT |
| Rapid Barcode Primer 08 | TTCAGGGAACAAACCAAGTTACGT |
| Rapid Barcode Primer 09 | AACTAGGCACAGCGAGTCTTGGTT |
| Rapid Barcode Primer 10 | AAGCGTTGAAACCTTTGTCCTCTC |
| Rapid Barcode Primer 11 | GTTTCATCTATCGGAGGGAATGGA |
| Rapid Barcode Primer 12 | GTTGAGTTACAAAGCACCGATCAG |
| Rapid Barcode Primer 13 | AGAACGACTTCCATACTCGTGTGA |
| Rapid Barcode Primer 14 | AACGAGTCTCTTGGGACCCATAGA |
| Rapid Barcode Primer 15 | AGGTCTACCTCGCTAACACCACTG |
| Rapid Barcode Primer 16 | CGTCAACTGACAGTGGTTCGTACT |
| Rapid Barcode Primer 17 | ACCCTCCAGGAAAGTACCTCTGAT |
| Rapid Barcode Primer 18 | CCAAACCCAACAACCTAGATAGGC |
| Rapid Barcode Primer 19 | GTTCCTCGTGCAGTGTCAAGAGAT |
| Rapid Barcode Primer 20 | TTGCGTCCTGTTACGAGAACTCAT |
| Rapid Barcode Primer 21 | GAGCCTCTCATTGTCCGTTCTCTA |
| Rapid Barcode Primer 22 | ACCACTGCCATGTATCAAAGTACG |
| Rapid Barcode Primer 23 | CTTACTACCCAGTGAACCTCCTCG |
| Rapid Barcode Primer 24 | GCATAGTTCTGCATGATGGGTTAG |
3. Library preparation
Materials
- Up to 24x single Salmonella colonies (2–3 mm in diameter)
- Fragmentation Mix (FRM)
- Rapid Barcode Primers (vial format (RLB01-24, at 10 µM) or plate format (RPB01-24, at 1 µM))
- EDTA (EDTA)
- AMPure XP Beads (AXP)
- Elution Buffer (EB)
- Rapid Adapter (RA)
- Adapter Buffer (ADB)
Consumables
- 1 M Tris Buffer, pH8 (e.g. Invitrogen™, AM9855G)
- LongAmp Hot Start Taq 2X Master Mix (NEB, M0533)
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- 1.5 ml Eppendorf DNA LoBind tubes
- 0.2 ml thin-walled PCR tubes
Equipment
- Inoculation loop or equivalent for picking your colony
- Ice bucket with ice
- Thermal cycler
- Hula mixer (gentle rotator mixer)
- Magnetic separation rack
- Microfuge
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- P2 pipette and tips
Optional equipment
- Microbiological Safety Cabinet II contamination level 3
Ensure pathogenic organisms and samples are handled in appropriate biosafety conditions.
Please adhere to the correct health and safety practices in accordance to your laboratory standards, and local rules and regulations.
This method should be performed in the appropriate containment level laboratory spaces (biosafety cabinets: BSC) until the sample tagmentation stage. Following sample tagmentation, cells are non-viable and can be used outside the biosafety cabinet.
Check your flow cell.
We recommend performing a flow cell check before starting your library prep to ensure you have a flow cell with enough pores for a good sequencing run.
See the flow cell check document for more information.
Thaw kit components at room temperature, spin down briefly using a microfuge and mix by pipetting as indicated by the Table below:
| Reagent | 1. Thaw at room temperature | 2. Briefly spin down | 3. Mix well by pipetting or vortexing |
|---|---|---|---|
| Rapid Barcode Primers (vial format (RLB01-24, at 10 µM) or plate format (RPB01-24, at 1 µM)) | ✓ | ✓ | Pipette |
| Fragmentation Mix (FRM) | Not frozen | ✓ | Pipette |
| Rapid Adapter (RA) | Not frozen | ✓ | Pipette |
| Adapter Buffer (ADB) | ✓ | ✓ | Vortex or Pipette |
| AMPure XP Beads (AXP) | ✓ | ✓ | Mix by pipetting or vortexing immediately before use |
| Elution Buffer (EB) | ✓ | ✓ | Vortex or Pipette |
| EDTA (EDTA) | ✓ | ✓ | Vortex or Pipette |
Note: Once thawed, keep all kit components on ice.
The wells of the barcoding plate are intended for single use only. Please ensure your barcode well is sealed before use, and do not reuse the barcode well once pierced/opened.
When picking a single colony, please ensure that no media is carried through as this can negatively impact the PCR yields.
Prepare each cultured cell colony as follows:
- For each colony/sample, add 50 µl 10 mM Tris buffer (pH 8) to a 1.5 ml Eppendorf DNA LoBind tube.
- Using an inoculation loop, transfer a single 2–3 mm colony from an overnight culture into a separate tube (from the step above).
- Resuspend each colony thoroughly by pipette mixing and briefly vortexing.
- Ensure your cell colony suspension is homogenous before proceeding.
In a separate 0.2 ml thin-walled PCR tube for each sample, mix the following:
| Reagent | Volume |
|---|---|
| DNA/colony suspension (from previous step) | 3 μl |
| Fragmentation Mix (FRM) | 1 μl |
| Total | 4 μl |
Mix thoroughly by pipetting the reaction.
In a thermal cycler, incubate the tube at 30°C for 2 minutes, then at 80°C for 2 minutes.
Sample tagmentation has taken place, cells are now non-viable and can be used outside the biosafety cabinet.
Please continue to adhere to the correct health and safety practices in accordance to your laboratory standards, and local rules and regulations.
Please ensure you are following the correct method for your SQK-RPB114.24 kit format.
In addition to the kit format change, we have lowered the concentration of the Rapid Barcode Primers in the plate format to 1 µM (down from 10 µM in the vial format) to facilitate automation usage and improve kit usability.
Please note, this will mean that the volumes used in the PCR reaction set up will vary depending on the kit format you have available.
For each sample, set up a PCR reaction as follows in the 0.2 ml thin-walled PCR tube containing the tagmented DNA:
Plate format with Rapid Barcode Primers at 1 µM
| Reagent | Volume |
|---|---|
| Tagmented DNA (from previous step) | 4 µl |
| Nuclease-free water | 11 µl |
| Rapid Barcode Primers (01-24, at 1 µM) | 10 µl |
| LongAmp Hot Start Taq 2X Master Mix | 25 µl |
| Total | 50 µl |
If the amount of input material is altered, the number of PCR cycles may need to be adjusted to produce the same yield.
Vial format with Rapid Barcode Primers at 10 µM
| Reagent | Volume |
|---|---|
| Tagmented DNA (from previous step) | 4 µl |
| Nuclease-free water | 20 µl |
| Rapid Barcode Primers (01-24, at 10 µM) | 1 µl |
| LongAmp Hot Start Taq 2X Master Mix | 25 µl |
| Total | 50 µl |
If the amount of input material is altered, the number of PCR cycles may need to be adjusted to produce the same yield.
Mix gently by flicking the tube, and spin down.
Amplify using the following cycling conditions:
| Cycle step | Temperature | Time | No. of cycles |
|---|---|---|---|
| Initial denaturation | 95°C | 3 mins | 1 |
| Denaturation Annealing Extension | 95°C 56°C 65°C | 15 sec 15 sec 6 min | 25* |
| Final extension | 65°C | 6 min | 1 |
| Hold | 4°C | ∞ |
*For more information on the PCR cycle number and externsion times, please visit our know-how document.
Add 4 µl of EDTA to each barcoded sample. Mix thoroughly by pipetting and spin down briefly.
EDTA is added at this step to stop the reaction.
Incubate for 5 minutes at room temperature.
Quantify 1 µl of each barcoded sample using a Qubit fluorometer (or equivalent) for QC check.
Note: The expected PCR yield for each sample is 15–45 ng/µl.
Pool 100 ng of each PCR barcoded sample in a clean 1.5 ml Eppendorf DNA LoBind tube.
Note: Please ensure you have quantified your samples prior to this step and take forward 100 ng of each of the samples for optimal barcode balancing.
Samples may vary in concentration following the barcoded PCR. Therefore, the volume of each barcoded sample added to the pool will be different.
Make up the volume of pooled samples to the nearest 100 µl using nuclease-free water.
For example:
- If the 100 ng pool of each sample results in a total of 88 µl, add 12 µl of nuclease-free water to make up the volume to 100 µl.
- If the 100 ng pool of each sample results in a total of 180 µl, add 20 µl of nuclease-free water to make up the volume to 200 µl.
Resuspend the AMPure XP Beads (AXP) by vortexing.
To the pool of barcoded samples, add a 0.6X volume ratio of resuspended AMPure XP Beads (AXP) and mix by pipetting:
| Volume of barcoded sample pool | 100 μl | 200 μl | 300 μl |
|---|---|---|---|
| Volume of AMPure XP Beads (AXP) | 60 μl | 120 μl | 180 μl |
Incubate the reaction on a hula mixer (rotator mixer) for 10 minutes at room temperature.
Prepare 2 ml of fresh 80% ethanol in nuclease-free water.
Briefly spin down the sample and pellet on a magnetic rack until the supernatant is clear and colourless. Keep the tube on the magnetic rack, and pipette off the supernatant.
Keep the tube on the magnet and wash the beads with 1 ml of freshly-prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Repeat the previous step.
Spin down and place the tube back on the magnet. Pipette off any residual ethanol. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend the pellet by pipetting in 15 µl Elution Buffer (EB). Spin down and incubate for 5 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless, for at least 1 minute.
Remove and retain 15 µl of eluate into a clean 1.5 ml Eppendorf DNA LoBind tube.
- Remove and retain the eluate which contains the DNA library in a clean 1.5 ml Eppendorf DNA LoBind tube.
- Dispose of the pelleted beads.
Quantify 1 µl of eluted sample using a Qubit fluorometer.
Transfer 250 ng of your eluted samples into a clean 1.5 ml Eppendorf DNA LoBind tube. Make up the volume to 11 µl with Elution Buffer (EB).
Note: If your final yield is below 250 ng, take forward 11 µl of eluted sample for the maxium flow cell loading amount possible.
Please note, if you underload the flow cell you might need to sequence for longer than stated in this method to achieve the recommended number of reads (at least 50k reads per barcode).
In a fresh 1.5 ml Eppendorf DNA LoBind tube, dilute the Rapid Adapter (RA) as follows and pipette mix:
| Reagent | Volume |
|---|---|
| Rapid Adapter (RA) | 1.5 μl |
| Adapter Buffer (ADB) | 3.5 μl |
| Total | 5 μl |
Add 1 µl of the diluted Rapid Adapter (RA) to the barcoded DNA.
Mix gently by flicking the tube, and spin down.
Incubate the reaction for 5 minutes at room temperature.
The prepared library is used for loading into the flow cell. Store the library on ice until ready to load.
4. Priming and loading the MinION and GridION Flow Cell
Materials
- Flow Cell Flush (FCF)
- Flow Cell Tether (FCT)
- Library Beads (LIB)
- Sequencing Buffer (SB)
Consumables
- MinION/GridION Flow Cell
- Bovine Serum Albumin (BSA) (50 mg/ml) (e.g. Invitrogen™ UltraPure™ BSA 50 mg/ml, AM2616)
- 1.5 ml Eppendorf DNA LoBind tubes
Equipment
- MinION or GridION device
- MinION/GridION Flow Cell Light Shield
- P1000 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
Please note, this kit is only compatible with R10.4.1 flow cells (FLO-MIN114).
Take the flow cell out of the fridge and leave it at room temperature for 20 minutes. This will improve visibility of the array during priming and sample loading.
Priming and loading a flow cell
We recommend all new users watch the 'Priming and loading your flow cell' video before your first run.
Thaw the Sequencing Buffer (SB), Library Beads (LIB), Flow Cell Tether (FCT) and Flow Cell Flush (FCF) at room temperature before mixing by vortexing. Then spin down and store on ice.
For optimal sequencing performance and improved output on MinION R10.4.1 flow cells (FLO-MIN114), add Bovine Serum Albumin (BSA) to the flow cell priming mix at a final concentration of 0.2 mg/ml.
Note: We do not recommend using any other albumin type (e.g. recombinant human serum albumin).
To prepare the flow cell priming mix with BSA, combine the following reagents in a fresh 1.5 ml Eppendorf DNA LoBind tube. Mix by inverting the tube and pipette mix at room temperature:
| Reagents | Volume per flow cell |
|---|---|
| Flow Cell Flush (FCF) | 1,170 µl |
| Bovine Serum Albumin (BSA) at 50 mg/ml | 5 µl |
| Flow Cell Tether (FCT) | 30 µl |
| Final total volume in tube | 1,205 µl |
Open the MinION or GridION device lid and slide the flow cell under the clip. Press down firmly on the priming port cover to ensure correct thermal and electrical contact.
Complete a flow cell check to assess the number of pores available before loading the library.
This step can be omitted if the flow cell has been checked previously.
See the flow cell check document for more information.
Slide the flow cell priming port cover clockwise to open the priming port.
Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µl, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the priming port
- Turn the wheel until the dial shows 220-230 µl, to draw back 20-30 µl, or until you can see a small volume of buffer entering the pipette tip
Note: Visually check that there is continuous buffer from the priming port across the sensor array.
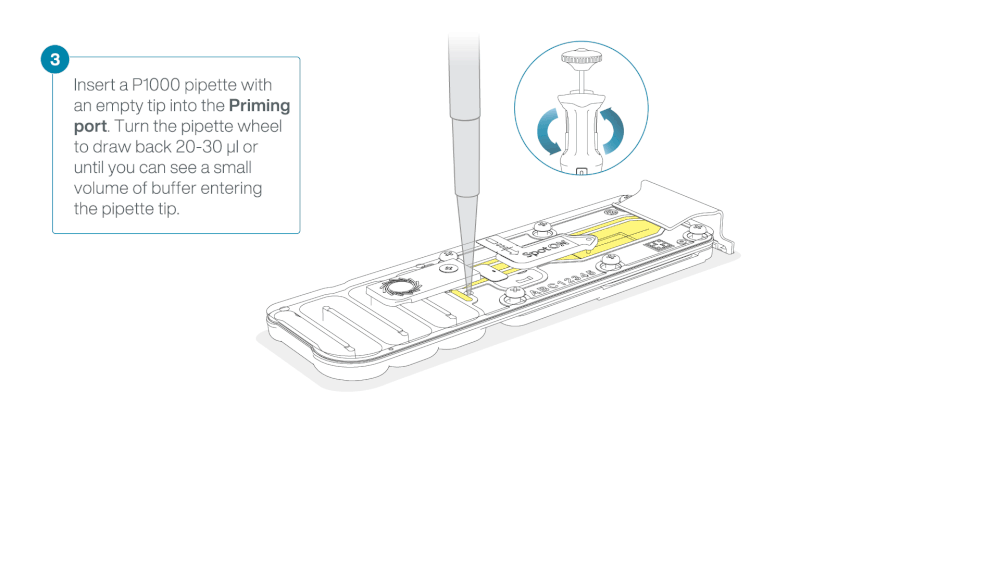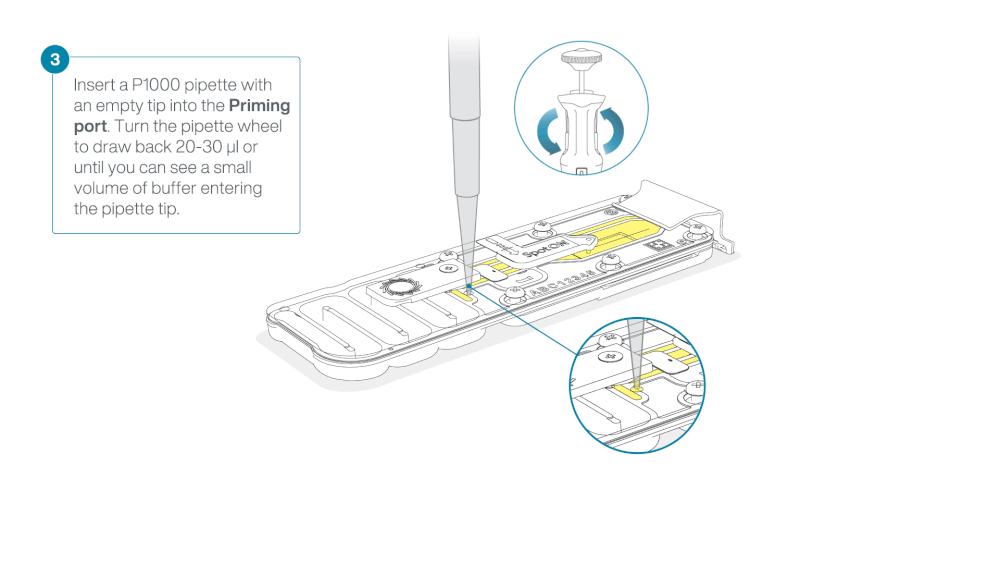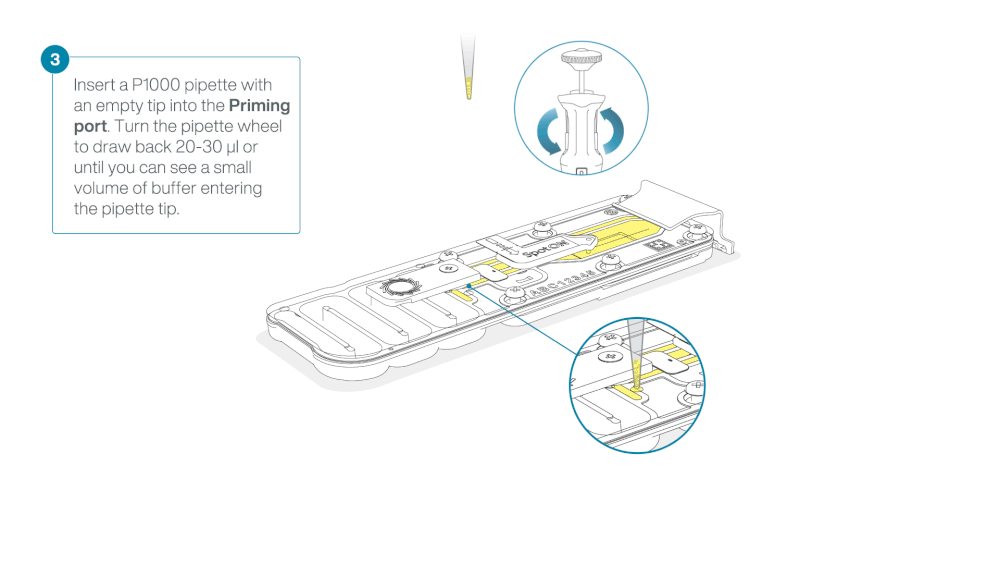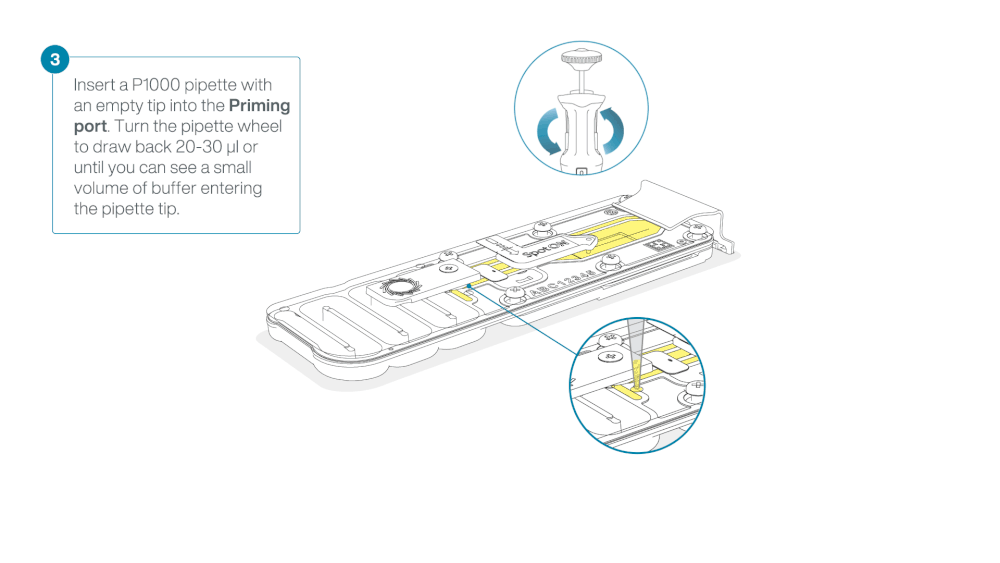
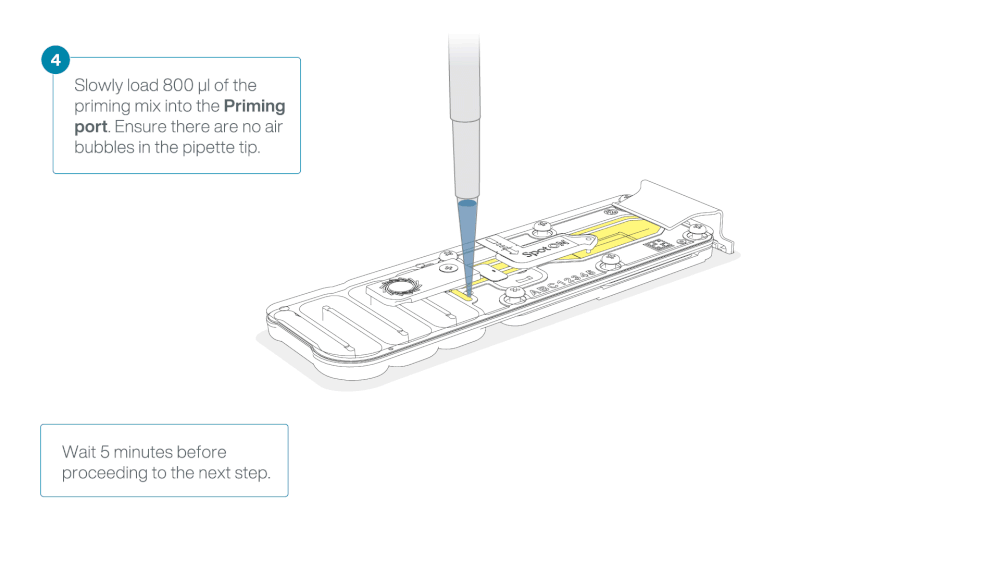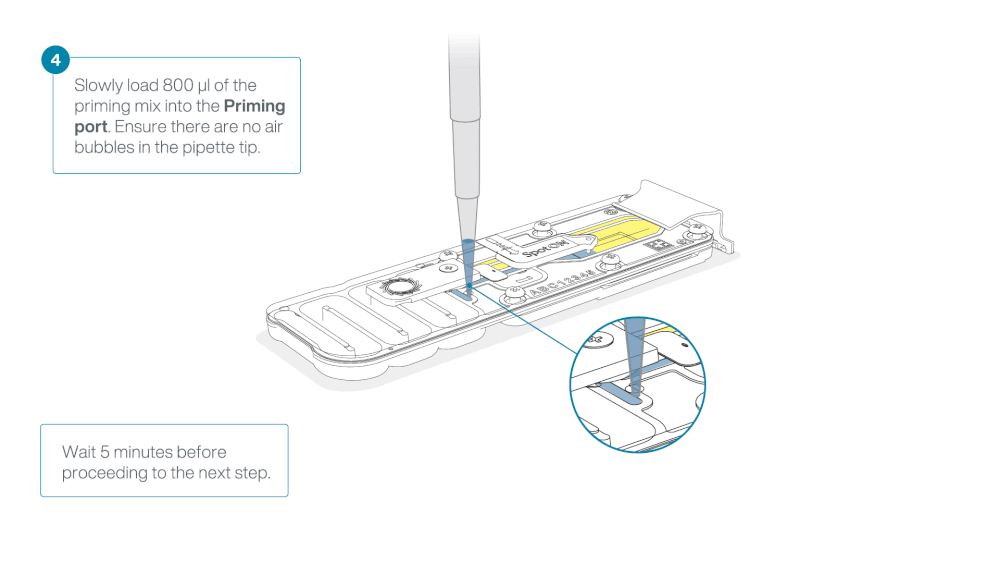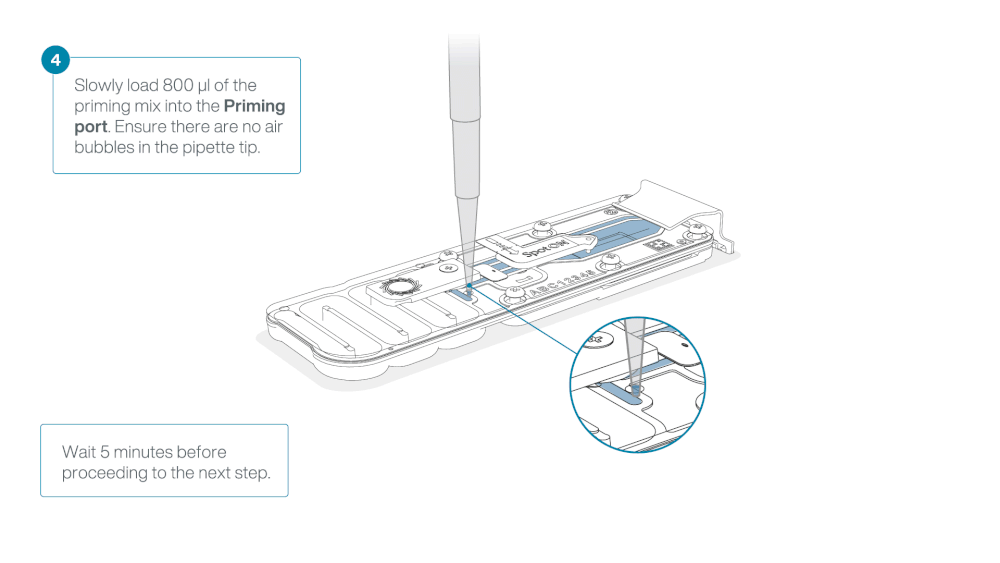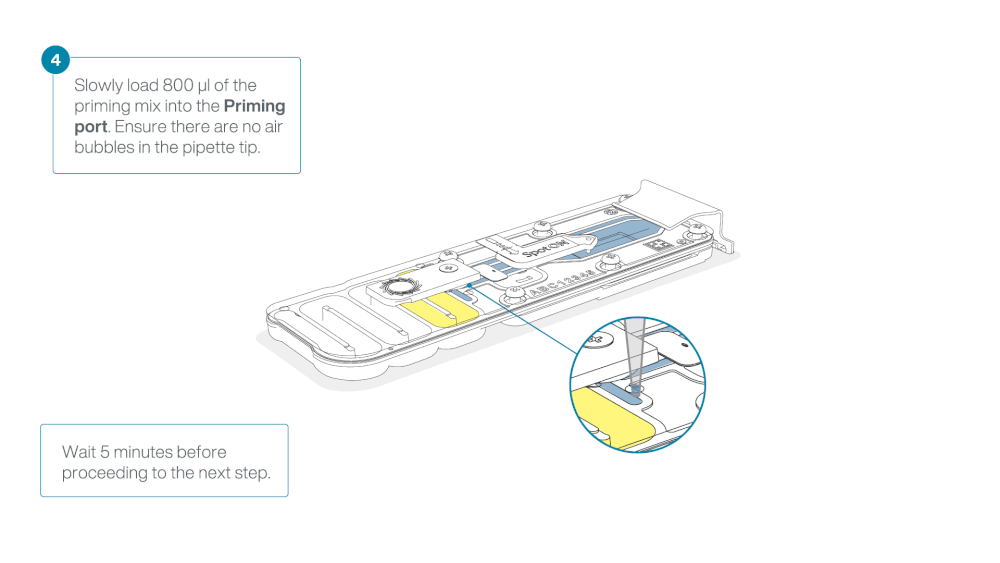
Load 800 µl of the priming mix into the flow cell via the priming port, avoiding the introduction of air bubbles. Wait for five minutes. During this time, prepare the library for loading by following the steps below.
Thoroughly mix the contents of the Library Beads (LIB) by pipetting.
The Library Beads (LIB) tube contains a suspension of beads. These beads settle very quickly. It is vital that they are mixed immediately before use.
We recommend using the Library Beads (LIB) for most sequencing experiments. However, the Library Solution (LIS) is available for more viscous libraries.
In a new 1.5 ml Eppendorf DNA LoBind tube, prepare the library for loading as follows:
| Reagent | Volume per flow cell |
|---|---|
| Sequencing Buffer (SB) | 37.5 µl |
| Library Beads (LIB) mixed immediately before use | 25.5 µl |
| DNA library | 12 µl |
| Total | 75 µl |
Complete the flow cell priming:
- Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
- Load 200 µl of the priming mix into the flow cell priming port (not the SpotON sample port), avoiding the introduction of air bubbles.

Mix the prepared library gently by pipetting up and down just prior to loading.
Add 75 μl of the prepared library to the flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.

Gently replace the SpotON sample port cover, making sure the bung enters the SpotON port and close the priming port.

For optimal sequencing output, install the light shield on your flow cell as soon as the library has been loaded.
We recommend leaving the light shield on the flow cell when library is loaded, including during any washing and reloading steps. The shield can be removed when the library has been removed from the flow cell.
Place the light shield onto the flow cell, as follows:
Carefully place the leading edge of the light shield against the clip. Note: Do not force the light shield underneath the clip.
Gently lower the light shield onto the flow cell. The light shield should sit around the SpotON cover, covering the entire top section of the flow cell.

The MinION Flow Cell Light Shield is not secured to the flow cell and careful handling is required after installation.
Close the device lid and set up a sequencing run on MinKNOW.
When a flow cell is inserted into the MinION Mk1D, the device lid will sit on top of the flow cell, leaving a small gap around the sides. This is normal and has no impact on the performance of the device.
Please refer to this FAQ regarding the device lid.

5. Data acquisition and basecalling
How to start sequencing
The MinKNOW software controls the sequencing device and carries out data acquisition and real-time basecalling. Please ensure MinKNOW is installed on your computer or device. Further instructions for setting up a sequencing run can be found in the MinKNOW protocol.
We recommend setting up a sequencing run on a MinION or GridION device using the basecalling and barcoding recommendations outlined below. All other parameters can be left to their default settings.
MinKNOW settings for Salmonella serotyping
For the fastest turnaround time and easiest analysis, basecalling should be performed live during the sequencing run using the High Accuracy (HAC) basecaller.
Below are the recommendations for MinKNOW settings:
Positions
Flow cell position: [user defined]
Experiment name: [user defined]
Flow cell type: FLO-MIN114
Sample ID: [user defined]
Kit
Kit selection: Rapid PCR Barcoding Kit (SQK-RPB114-24)
Run configuration
Sequencing and analysis
Basecalling: On [default]
Modified bases: Off
Model: High-accuracy basecalling (HAC) [default]
Barcoding: On [default]
Trim barcodes: Off [default]
Barcode both ends: Off [default]
Custom barcodes selection: Off [default]
Alignment: Off [default]
Adaptive sampling: Off [default]
Advanced options
Active channel selection: On [default]
Time between pore scans: 1.5 [default]
Reserve pores: On [default]
Data targets
Sequencing time will vary according to the number of samples being sequenced. The aim is to generate 50k reads per sample/barcode.
Run limit: Set according to number of samples:
| Number of samples/barcodes | Recommended sequencing time |
|---|---|
| 6 | 4 hours |
| 12 | 8 hours |
| 24 | 12 hours |
Output
Output format
.POD5: On [default]
.FASTQ: On [default]
.BAM: On
Filtering: On [default]
Qscore: 9 [default]
Minimum read length: 200 bp [default]
6. Downstream analysis
Post-basecalling analysis
We recommend performing downstream analysis using EPI2ME which facilitates bioinformatic analyses by allowing users to run Nextflow workflows in a desktop application. EPI2ME maintains a collection of bioinformatic workflows which are curated and actively maintained by experts in long-read sequence analysis.
Further information about the available EPI2ME workflows are available here, along with the Quick Start Guide to start your first bioinformatic workflow.
For the accurate and reliable assembly or alignment of you sample genomes, we recommend using the wf-bacterial-genomes workflow. At its core, the workflow is an efficient assembly pipeline which also polishes genomes using Medaka.
While running the workflow using the default parameters, use the ‘Isolates’ mode to produce high quality genome assemblies, perform additional analyses designed to increase genome quality and aid genome interrogation for common pathogens in the clinical and food safety fields. ‘Isolates’ analyses includes MLST(7-gene), species confirmation and AMR prediction in addition to sample-specific reports.
Note: You can also run this workflow through the command line. However, we only recommend this option for experienced users. For more information, please visit the wf-bacterial-genomes page on GitHub and the know-how document.
Please ensure your device meets the minimum compute requirements to run this workflow.
Compute requirements:
| CPUs | Memory | |
|---|---|---|
| Minimum requirements: | 8 | 32 GB |
| Recommended requirements: | 16 | 64 GB |
For more information visit the EPI2ME wf-bacterial-genomes documentation.
Open the EPI2ME app using the desktop shortcut.
On the landing page, open the workflow tab on the left-hand sidebar.

Navigate to the Available Workflows tab and click on wf-bacterial-genomes option.

Click install.

Navigate to the Installed tab and click on the installed wf-bacterial-genomes workflow.

If the workflow was already installed, check for updates by clicking 'Update workflow'.
Ensure you are using v1.3.0 or newer of the workflow. We recommend running the latest version of our workflows for the best results.


Click on Run this workflow to open the launch wizard.

Set up your run by uploading your FASTQ file in the Input Options. We recommend keeping the default settings for the other parameter options.

In Sample Options, provide a sample sheet to match the sample folder names (e.g. barcode01) to a sample name alias to be used in output reports.
You may specify any number of sample (barcode) folders in your sample sheet.

To enable the Isolates mode, tick the Isolates checkbox.

In Advanced Options unselect Prokka annotation.
Optional setting: Set Flye genome size to 5000000, and Flye asm coverage to 50 (these settings are optional but will speed up run time).

In Miscellaneous Options, set number of CPU threads.
Navigate to the 'Nextflow configuration' tab to assign a 'Run name' to your analysis as an identifier.

Click Launch workflow.
Ensure all parameter options have green ticks.

Once the workflow finishes, a report will be produced.
Test data is available on Github to test the wf-bacterial-genomes workflow.
The test data can be found in the following repository: wf-bacterial-genomes test data
7. Flow cell reuse and returns
Materials
- Flow Cell Wash Kit (EXP-WSH004)
After your sequencing experiment is complete, if you would like to reuse the flow cell, please follow the Flow Cell Wash Kit protocol and store the washed flow cell at +2°C to +8°C.
The Flow Cell Wash Kit protocol is available on the Nanopore Community.
We recommend you to wash the flow cell as soon as possible after you stop the run. However, if this is not possible, leave the flow cell on the device and wash it the next day.
Alternatively, follow the returns procedure to send the flow cell back to Oxford Nanopore.
Instructions for returning flow cells can be found here.
If you encounter issues or have questions about your sequencing experiment, please refer to the Troubleshooting Guide that can be found in this protocol.
8. Issues during DNA extraction and library preparation
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Low sample quality
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) | The DNA extraction method does not provide the required purity | The effects of contaminants are shown in the Contaminants document. Please try an alternative extraction method that does not result in contaminant carryover. Consider performing an additional SPRI clean-up step. |
Low DNA recovery after AMPure bead clean-up
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low recovery | DNA loss due to a lower than intended AMPure beads-to-sample ratio | 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample. 2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up. |
| Low recovery | DNA fragments are shorter than expected | The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 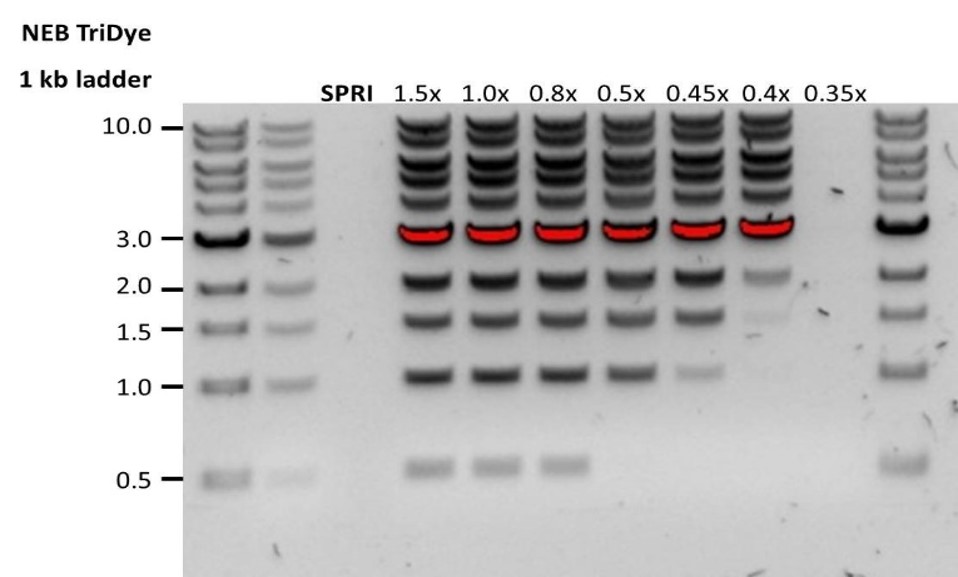 |
| Low recovery after end-prep | The wash step used ethanol <80% | DNA will be eluted from the beads when using ethanol <80%. Make sure to use the correct percentage. |
9. Issues during the sequencing run using a Rapid-based sequencing kit
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Fewer pores at the start of sequencing than after Flow Cell Check
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | An air bubble was introduced into the nanopore array | After the Flow Cell Check it is essential to remove any air bubbles near the priming port before priming the flow cell. If not removed, the air bubble can travel to the nanopore array and irreversibly damage the nanopores that have been exposed to air. The best practice to prevent this from happening is demonstrated in this video. |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | The flow cell is not correctly inserted into the device | Stop the sequencing run, remove the flow cell from the sequencing device and insert it again, checking that the flow cell is firmly seated in the device and that it has reached the target temperature. If applicable, try a different position on the device (GridION/PromethION). |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | Contaminations in the library damaged or blocked the pores | The pore count during the Flow Cell Check is performed using the QC DNA molecules present in the flow cell storage buffer. At the start of sequencing, the library itself is used to estimate the number of active pores. Because of this, variability of about 10% in the number of pores is expected. A significantly lower pore count reported at the start of sequencing can be due to contaminants in the library that have damaged the membranes or blocked the pores. Alternative DNA/RNA extraction or purification methods may be needed to improve the purity of the input material. The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
MinKNOW script failed
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Script failed" | Restart the computer and then restart MinKNOW. If the issue persists, please collect the MinKNOW log files and contact Technical Support. If you do not have another sequencing device available, we recommend storing the flow cell and the loaded library at 4°C and contact Technical Support for further storage guidance. |
Pore occupancy below 40%
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Pore occupancy <40% | Not enough library was loaded on the flow cell | 10–50 fmol of good quality library can be loaded on to a MinION Mk1B/GridION flow cell. Please quantify the library before loading and calculate mols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to pmol" |
| Pore occupancy close to 0 | The Rapid PCR Barcoding Kit V14 was used, and sequencing adapters did not attach to the DNA | Make sure to closely follow the protocol and use the correct volumes and incubation temperatures. A Lambda control library can be prepared to test the integrity of reagents. |
| Pore occupancy close to 0 | No tether on the flow cell | Tethers are added during flow cell priming (FCT tube). Make sure FCT was added to FCF before priming. |
Shorter than expected read length
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Shorter than expected read length | Unwanted fragmentation of DNA sample | Read length reflects input DNA fragment length. Input DNA can be fragmented during extraction and library prep. 1. Please review the Extraction Methods in the Nanopore Community for best practice for extraction. 2. Visualise the input DNA fragment length distribution on an agarose gel before proceeding to the library prep. 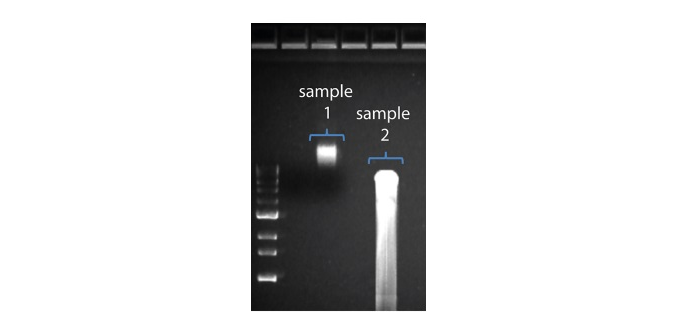 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented. In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.3. During library prep, avoid pipetting and vortexing when mixing reagents. Flicking or inverting the tube is sufficient. |
Large proportion of unavailable pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
Large proportion of unavailable pores (shown as blue in the channels panel and pore activity plot) 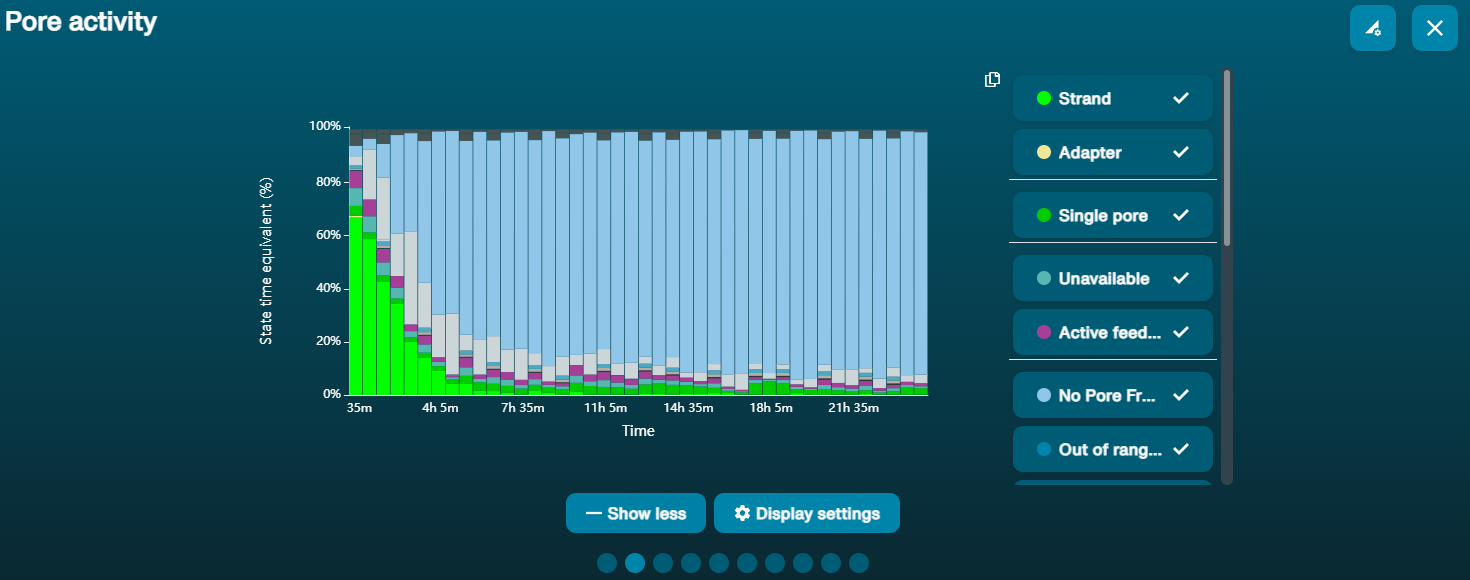 The pore activity plot above shows an increasing proportion of "unavailable" pores over time. The pore activity plot above shows an increasing proportion of "unavailable" pores over time. | Contaminants are present in the sample | Some contaminants can be cleared from the pores by the unblocking function built into MinKNOW. If this is successful, the pore status will change to "sequencing pore". If the portion of unavailable pores stays large or increases: 1. A nuclease flush using the Flow Cell Wash Kit (EXP-WSH004) can be performed, or 2. Run several cycles of PCR to try and dilute any contaminants that may be causing problems. |
Large proportion of inactive pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Large proportion of inactive/unavailable pores (shown as light blue in the channels panel and pore activity plot. Pores or membranes are irreversibly damaged) | Air bubbles have been introduced into the flow cell | Air bubbles introduced through flow cell priming and library loading can irreversibly damage the pores. Watch the Priming and loading your flow cell video for best practice |
| Large proportion of inactive/unavailable pores | Certain compounds co-purified with DNA | Known compounds include polysaccharides. Clean-up using the QIAGEN PowerClean Pro kit. |
| Large proportion of inactive/unavailable pores | Contaminants are present in the sample | The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
Temperature fluctuation
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Temperature fluctuation | The flow cell has lost contact with the device | Check that there is a heat pad covering the metal plate on the back of the flow cell. Re-insert the flow cell and press it down to make sure the connector pins are firmly in contact with the device. If the problem persists, please contact Technical Services. |
Failed to reach target temperature
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Failed to reach target temperature" | The instrument was placed in a location that is colder than normal room temperature, or a location with poor ventilation (which leads to the flow cells overheating) | MinKNOW has a default timeframe for the flow cell to reach the target temperature. Once the timeframe is exceeded, an error message will appear and the sequencing experiment will continue. However, sequencing at an incorrect temperature may lead to a decrease in throughput and lower q-scores. Please adjust the location of the sequencing device to ensure that it is placed at room temperature with good ventilation, then re-start the process in MinKNOW. Please refer to this link for more information on MinION temperature control. |







