Ligation sequencing DNA V14 - automated Tecan DreamPrep NGS (SQK-LSK114-XL) (DTD_9181_v114_revL_02Oct2025)
GridION: Protocol
Ligation sequencing DNA V14 - automated Tecan DreamPrep NGS (SQK-LSK114-XL) V DTD_9181_v114_revL_02Oct2025
This protocol:
- Uses genomic DNA
- Has a hands-on setup time ~30 minutes
- Has a automated run time ~2 hours 40 minutes
- Has a throughput: process 8–96 samples for library loading
- Is compatible with R10.4.1 Flow Cells
For Research Use Only
FOR RESEARCH USE ONLY.
Contents
Introduction to the protocol
Automated library preparation
- 3. Library preparation
- 4. Priming and loading the MinION/GridION Flow Cell
- 5. Unloading the Tecan DreamPrep NGS worktable
Sequencing and data analysis
Troubleshooting
Disclaimers
Overview
This protocol:
- Uses genomic DNA
- Has a hands-on setup time ~30 minutes
- Has a automated run time ~2 hours 40 minutes
- Has a throughput: process 8–96 samples for library loading
- Is compatible with R10.4.1 Flow Cells
For Research Use Only
1. Overview of the protocol
This protocol describes the automated workflow using the Ligation Sequencing Kit XL (SQK-LSK114-XL).
All images and information reflect the use of SQK-LSK114-XL.
For more information on compatibilities and performing an automated library preparation with previous iterations of the Ligation Sequencing Kit, please contact us on our website.
Ligation Sequencing Kit XL V14 features
This kit is recommended for:
- Processing multiple samples simultaneously, either with a multichannel pipette or a liquid-handling robot
- Achieving raw read sequencing modal accuracy of Q20+ (99%) or above
- Controlling read length
- Upstream processes such as size selection, whole genome amplification, or enrichment for long reads
Optional fragmentation and size selection
By default, the protocol contains no DNA fragmentation step, however in some cases it may be advantageous to fragment your sample. For example, when working with lower amounts of input gDNA (100 ng – 500 ng), fragmentation will increase the number of DNA molecules and therefore increase throughput. Instructions are available in the DNA Fragmentation section of Extraction methods.
Additionally, we offer several options for size-selecting your DNA sample to enrich for long fragments - instructions are available in the Size Selection section of Extraction methods.
Introduction to the automated Ligation Sequencing protocol for DNA
This protocol describes how to carry out sequencing of a DNA sample using the Ligation Sequencing Kit XL (SQK-LSK114-XL).
We have developed this automated protocol on the Tecan® DreamPrep® NGS liquid handling robot. This library preparation protocol is fully automated, except for the sample quantification steps and loading of the system.
Please note that this method is intended for research use only.
It is highly recommended that a Lambda control experiment is completed first to become familiar with the technology.
Steps in the sequencing workflow:
Prepare for your experiment
Before starting - Manual steps:
- Extract your DNA and check its length, quantity and purity. The quality checks performed during the protocol are essential in ensuring experimental success.
- Ensure you have your sequencing kit, the correct equipment, primed liquid-handling robot and third-party reagents.
- Download the software for acquiring and analysing your data.
- Check your flow cells to ensure they have enough pores for a good sequencing run.
Prepare your library
Automated steps:
- Repair the DNA and prepare the DNA ends for adapter attachment.
- Attach sequencing adapters supplied in the kit to the DNA ends.
Manual steps:
- Quantify your DNA library as a quality control.
- Prime the flow cell and load your DNA library into the flow cell.
#### Overview of library preparation workflow:
Note: Timings are dependent on number of samples and include hands on time, such as deck loading and sample quantification.
Sequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software which will collect raw data from the device and convert it into basecalled reads.
Timings
Note: Timings are approximate and subject to change with updates.
| Process | X8 samples | X24 samples | X48 samples | X96 samples | Hands-on time |
|---|---|---|---|---|---|
| Deck set-up and master mix preparation | ~30 minutes | ||||
| Automated DNA repair, end-prep and clean-up | 1 hour | 1 hour | 1 hour 5 minutes | 1 hour 10 minutes | |
| Automated adapter ligation and clean-up | 1 hour 30 minutes | 1 hour 30 minutes | 1 hour 35 minutes | 1 hour 35 minutes | |
| Quantification | ~10 minutes | ||||
| Total | 2 hours 30 minutes | 2 hours 30 minutes | 2 hours 40 minutes | 2 hours 45 minutes | ~40 minutes |
Sequencing run set-up and flow cell loading timings are variable depending on the number of samples and user experience.
Compatibility of this protocol
This protocol should only be used in combination with:
- Ligation Sequencing Kit XL V14 (SQK-LSK114-XL)
- R10.4.1 flow cells (FLO-MIN114)
- Flow Cell Wash Kit XL (EXP-WSH004-XL)
- MinION Mk1D - MinION Mk1D IT requirements document
- GridION - GridION IT requirements document
2. Equipment and consumables
Materials
- 1 µg (or 100-200 fmol) high molecular weight genomic DNA
- OR >100 ng high molecular weight genomic DNA if performing DNA fragmentation
- Ligation Sequencing Kit XL V14 (SQK-LSK114-XL)
Consumables
- NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing (NEB, E7180S or E7180L). Alternatively, you can use the NEBNext® products below:
- NEBNext FFPE Repair Mix (NEB, M6630)
- NEBNext Ultra II End repair/dA-tailing Module (NEB, E7546)
- NEBNext Quick Ligation Module (NEB, E6056)
- Agencourt AMPure XP beads (Beckman Coulter™, A63881)
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Bovine Serum Albumin (BSA) (50 mg/ml) (e.g. Invitrogen™ UltraPure™ BSA 50 mg/ml, AM2616)
- Hard-Shell® 96-Well PCR Plates, low profile, thin wall, skirted, red/clear (Bio-Rad™, HSP9611)
- Abgene™ 96 Well 1.2 ml Polypropylene Deepwell Storage Plate (ThermoFisher Scientific, AB1127)
- 1000 µl Disposable Conductive Tips - Liquid Handling Flexible Channel Arm - Filtered, Pure, ANSI/SLAS-format box (same as SBS) (Tecan, 30057817)
- 200 µl Disposable Conductive Tips - Liquid Handling Flexible Channel Arm - Filtered, Pure, ANSI/SLAS-format box (same as SBS) (Tecan, 30057815)
- 50 µl Disposable Conductive Tips - Liquid Handling Flexible Channel Arm - Filtered, Pure, ANSI/SLAS-format box (same as SBS) (Tecan, 30057813)
- Small SBS Box to place conductive tips & refill, compatible with 10 µl, 50 µl, 200 µl tips (Tecan, 30058506)
- Big SBS Box to place conductive tips & refill, compatible with 1000 µl tips (Tecan, 30058507)
- 150 µl Disposable Tips - MultiChannel Arm™ 384/96 - Filtered, Sterile, Single Stack (Tecan, 30038618)
- 50 µl Disposable Tips - MultiChannel Arm 384/96 - Filtered, Sterile, Single Stack (Tecan, 30038608)
- 100 ml disposable trough (Tecan, 10613049)
- 25 ml disposable trough (Tecan, 30055743)
- Arched Auto-Sealing Lids with Wide Tabs for PCR Plates (Bio-Rad™, MSL2032 or equivalent)
- Screw Cap Micro tube 2 ml, sterile (Sarstedt™, 72.694.321)
- 1.5 ml Eppendorf DNA LoBind tubes
- 2 ml Eppendorf DNA LoBind tubes
- 5 ml Eppendorf DNA LoBind tubes
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Low-lint scientific wipes (e.g. Kimberly-Clark™, 7552 or equivalent)
- Double distilled water (ddH2O) (e.g. ThermoFisher, 11983084)
- Freshly prepared ≥80% ethanol in nuclease-free or double distilled water, for cleaning
- ~10% Bleach (or equivalent): Thermo Scientific Alfa Aesar Sodium hypochlorite, 11-15% available chlorine, (e.g. ThermoFisher, 15429019)
Equipment
- Ice bucket with ice
- Vortex mixer
- Microplate centrifuge
- Tecan DreamPrep NGS workstation with full configuration
Optional equipment
- Agilent Bioanalyzer (or equivalent)
- Qubit fluorometer plate reader (or equivalent for QC check)
Adjust your sample input quantity depending on your initial DNA sample length:
| Fragment library length | Starting input |
|---|---|
| Very short (<1 kb) | 200 fmol |
| Short (1-10 kb) | 100–200 fmol |
| Long (>10 kb) | 1 µg |
For more information on sample input and flow cell loading amounts for our ligation sequencing protocols please visit our know-how document.
Input DNA
How to QC your input DNA
It is important to use a plate reader to ensure the input DNA meets the quantity and quality requirements. Using too little or too much DNA, or DNA of poor quality (e.g. highly fragmented or containing RNA or chemical contaminants) can affect your library preparation.
For instructions on how to perform quality control of your DNA sample, please read the Input DNA/RNA QC protocol.
Tecan DreamPrep NGS
This method has been tested and validated using the Tecan DreamPrep NGS including an on deck thermal cycler (ODTC). An option to not use the ODTC is available in the method. This protocol will require installation by a Tecan Application Specialist, please contact your Tecan representative for further details.
Tecan DreamPrep NGS worktable layout
After having the system installed according to specifications, users will need to load the automated workstation before running the protocol.
Please always refer to the latest instructions on how to load the worktable, which are displayed and explained in the touchscreen display (with TouchTools™).
Examples of how to load the system are displayed from steps 11 to 20, which can be found in the Library preparation section of this protocol.
Please also find below the reference images for the workbench layouts:

Trough mounting sites:

Hotel sites:

NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing
For customers new to nanopore sequencing, we recommend buying the NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing (NEB, E7180S or E7180L), which contains all the NEB reagents needed for use with the Ligation Sequencing Kit.
Please note, for our amplicon protocols, NEBNext FFPE DNA Repair Mix and NEBNext FFPE DNA Repair Buffer are not required.
Third-party reagents
We have validated and recommend the use of all the third-party reagents used in this protocol. Alternatives have not been tested by Oxford Nanopore Technologies.
For all third-party reagents, we recommend following the manufacturer's instructions to prepare the reagents for use.
Check your flow cell
We highly recommend that you check the number of pores in your flow cell prior to starting a sequencing experiment. This should be done within 12 weeks of purchasing your MinION/GridION/PromethION Flow Cells. Oxford Nanopore Technologies will replace any unused flow cell with fewer than the number of pores listed in the Table below, when the result is reported within two days of performing the flow cell check, and when the storage recommendations have been followed. To do the flow cell check, please follow the instructions in the Flow Cell Check document.
| Flow cell | Minimum number of active pores covered by warranty |
|---|---|
| MinION/GridION Flow Cell | 800 |
| PromethION Flow Cell | 5000 |
We strongly recommend using the Ligation Buffer (LNB) supplied in the Ligation Sequencing Kit V14 rather than any third-party ligase buffers to ensure high ligation efficiency of the Ligation Adapter (LA).
Ligation Adapter (LA) included in this kit and protocol is not interchangeable with other sequencing adapters.
Consumables and reagent quantities required:
| Consumables | X8 samples | X24 samples | X48 samples | X96 samples |
|---|---|---|---|---|
| Tecan 1000 µl Flexible Channel Arm filter tips | 49 tips | 50 tips | 51 tips | 53 tips |
| Tecan 200 µl Flexible Channel Arm filter tips | 33 tips | 32 tips | 32 tips | 32 tips |
| Tecan 50 µl Flexible Channel Arm filter tips | 16 tips | 16 tips | 16 tips | 16 tips |
| Tecan 150 µl MultiChannel Arm 384/96 filter tips | 1 box | 3 boxes | 5 boxes | 10 boxes |
| Tecan 50 µl MultiChannel Arm 384/96 filter tips | 1 box | 2 boxes | 3 boxes | 5 boxes |
| Bio-Rad Hard-Shell® 96-Well PCR Plate | 8 | 8 | 8 | 8 |
| Abgene™ 96 Well 1.2mL Polypropylene Deepwell Storage Plate | 1 | 1 | 1 | 1 |
| Tecan 25 ml disposable trough | 4 | 4 | 4 | 3 |
| Tecan 100 ml disposable trough | 1 | 1 | 1 | 2 |
| ODTC plate lid | 1 | 1 | 1 | 1 |
| Sarstedt Inc Screw Cap Micro tube 2 ml | 2 | 2 | 3 | 5 |
| Reagents/kits | X8 samples | X24 samples | X48 samples | X96 samples |
|---|---|---|---|---|
| 80% ethanol | 10 ml | 16 ml | 24 ml | 42 ml |
| Nuclease-free water | 3.5 ml | 4.6 ml | 6.3 ml | 9.5 ml |
| AMPure XP Beads | 3 ml | 4.9 ml | 7.8 ml | 13.5 ml |
| Ligation Sequencing Kit XL V14 (SQK-LSK114-XL) | 1 kit | 1 kit | 1 kit | 2 kits |
| NEBNext Companion Module for Oxford Nanopore Technologies Ligation Sequencing (Cat# E7180L) | 1 kit | 1 kit | 1 kit | 1 kit |
| NEBNext Companion Module for Oxford Nanopore Technologies Ligation Sequencing (Cat# E7180S) | - | - | - | 1 kit |
| Alternative to the NEBNext Companion Module, individual reagents can be bought as seen below: | - | - | - | - |
| NEBNext FFPE DNA Repair Mix (Cat# M6630L) | 1 kit | 1 kit | 1 kit | 2 kits |
| NEBNext Ultra II End Repair/dA-Tailing Module (Cat# E7546L) | 1 kit | 1 kit | 1 kits | 2 kits |
| NEBNext Quick Ligation Module (Cat# E6056L) | 1 kit | 1 kit | 1 kit | 2 kits |
| NEBNext Quick Ligation Module (Cat# E6056S) | - | - | 1 kit | 1 kit |
| Note: These are the number of kits required for one run through for the selected number of samples. |
Ligation Sequencing Kit XL V14 (SQK-LSK114-XL) contents
| Name | Acronym | Vial colour | Number of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| DNA Control Strand | DCS | Yellow | 1 | 100 |
| Ligation Adapter | LA | Green | 1 | 320 |
| Ligation Buffer | LNB | White | 1 | 1,500 |
| Elution Buffer | EB | White cap, black strip label | 1 | 10,000 |
| Long Fragment Buffer | LFB | White cap, orange strip label | 2 | 20,000 |
| Short Fragment Buffer | SFB | White cap, blue strip label | 2 | 25,000 |
| Library Beads | LIB | Pink | 2 | 1,800 |
| Library Solution | LIS | White cap, pink label | 2 | 1,800 |
| Sequencing Buffer | SB | Red | 3 | 1,700 |
| Flow Cell Flush | FCF | Clear | 4 | 15,500 |
| Flow Cell Tether | FCT | Purple | 1 | 1,600 |
Note: The DNA Control Sample (DCS) is a 3.6 kb standard amplicon mapping the 3' end of the Lambda genome.
3. Library preparation
Materials
- 1 µg (or 100-200 fmol) high molecular weight genomic DNA
- Ligation Adapter (LA)
- Ligation Buffer (LNB)
- Long Fragment Buffer (LFB)
- S Fragment Buffer (SFB)
- Elution Buffer from the Oxford Nanopore kit (EB)
Consumables
- NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing (NEB, E7180S or E7180L). Alternatively, you can use the NEBNext® products below:
- NEBNext® FFPE DNA Repair Mix (NEB, M6630)
- NEBNext® Ultra™ II End Repair/dA-Tailing Module (NEB, E7546)
- NEBNext Quick Ligation Module (NEB, E6056)
- Agencourt AMPure XP beads (Beckman Coulter™, A63881)
- Nuclease-free water (e.g. ThermoFisher, AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Hard-Shell® 96-Well PCR Plates, low profile, thin wall, skirted, red/clear (Bio-Rad™, HSP9611)
- Abgene™ 96 Well 1.2 ml Polypropylene Deepwell Storage Plate (ThermoFisher Scientific, AB1127)
- 5 ml Eppendorf DNA LoBind tubes
- 2 ml Eppendorf DNA LoBind tubes
- 1.5 ml Eppendorf DNA LoBind tubes
- 1000 µl Disposable Conductive Tips - Liquid Handling Flexible Channel Arm - Filtered, Pure, ANSI/SLAS-format box (same as SBS) (Tecan, 30057817)
- 200 µl Disposable Conductive Tips - Liquid Handling Flexible Channel Arm - Filtered, Pure, ANSI/SLAS-format box (same as SBS) (Tecan, 30057815)
- 50 µl Disposable Conductive Tips - Liquid Handling Flexible Channel Arm - Filtered, Pure, ANSI/SLAS-format box (same as SBS) (Tecan, 30057813)
- Small SBS Box to place conductive tips & refill, compatible with 10 µl, 50 µl, 200 µl tips (Tecan, 30058506)
- Big SBS Box to place conductive tips & refill, compatible with 1000 µl tips (Tecan, 30058507)
- 150 µl Disposable Tips - MultiChannel Arm™ 384/96 - Filtered, Sterile, Single Stack (Tecan, 30038618)
- 50 µl Disposable Tips - MultiChannel Arm 384/96 - Filtered, Sterile, Single Stack (Tecan, 30038608)
- 100 ml disposable trough (Tecan, 10613049)
- 25 ml disposable trough (Tecan, 30055743)
- Screw Cap Micro tube 2 ml, sterile (Sarstedt™, 72.694.321)
Equipment
- Ice bucket with ice
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P10 pipette and tips
- Microplate centrifuge
- Vortex mixer
Optional equipment
- Qubit fluorometer plate reader (or equivalent for QC check)
Please note, all screenshots are representative of X96 samples.
Optional fragmentation and size selection
By default, the protocol contains no DNA fragmentation step, however in some cases it may be advantageous to fragment your sample. For example, when working with lower amounts of input gDNA (100 ng – 500 ng), fragmentation will increase the number of DNA molecules and therefore increase throughput. Instructions are available in the DNA Fragmentation section of Extraction methods.
Additionally, we offer several options for size-selecting your DNA sample to enrich for long fragments - instructions are available in the Size Selection section of Extraction methods.
Prepare the NEBNext FFPE DNA Repair Mix and NEBNext Ultra II End Repair / dA-tailing Module reagents in accordance with manufacturer’s instructions, and place on ice.
For optimal performance, NEB recommend the following:
Thaw all reagents on ice.
Flick and/or invert the reagent tubes to ensure they are well mixed.
Note: Do not vortex the FFPE DNA Repair Mix or Ultra II End Prep Enzyme Mix.Always spin down tubes before opening for the first time each day.
The Ultra II End Prep Buffer and FFPE DNA Repair Buffer may have a little precipitate. Allow the mixtures to come to room temperature and pipette the buffer up and down several times to break up the precipitate, followed by vortexing the tube for 30 seconds to solubilise any precipitate.
Note: It is important the buffers are mixed well by vortexing.The FFPE DNA Repair Buffer may have a yellow tinge and is fine to use if yellow.
Switch on the Tecan DreamPrep NGS robot and open the Fluent Control software on the computer. Follow the recommended specifications to initiate the DreamPrep NGS.

Perform the 'Daily System Care' method to prepare the instrument before the first run of the day.
Users will have access to the 'Main screen' of TouchTools™, which allows interaction with the DreamPrep NGS system. Select 'Method Starter'.

In the 'Method Starter' folder, select the Ligation sequencing program and click 'Ok'.
Click the 'start button' in the middle of the screen to start the run.

During the run set-up on the Tecan DreamPrep NGS, the workstation will perform automated checks and set-up for the ODTC.
During the run set-up, messages will pop-up regarding the ODTC set-up.

Allow these to complete and continue your run set-up as normal.
When you see the 'Welcome to the ONT Ligation Sequencing protocol by Tecan' page, click 'Continue'.

Set the 'User defined variables' and click on 'Next page' to proceed.

Note: Any number of samples between 8-96 can be processed using this protocol.
Set the 'User inputs' and click on 'Next page' to proceed.
- Please note that if removing the sample plate for off-deck storage, user interaction is required approximately 10 minutes after starting the run.
- We recommend room temperature for the majority of users. However, 37°C can be beneficial for recovery of longer DNA strands.
- The MinION option is valid for both the MinION and GridION device.

Reminder: If selecting 'YES' on 'Store Sample plate off-deck after use', user interaction will be required 10 minutes after starting the run. Click 'Next Page' to proceed.

Select if you would like to maintain the standard LSK volumes and timers as recommended by ONT and click on 'Next page' to proceed.

Note: Please note only the ONT recommended settings have been verified.
We highly recommend using default settings developed by Oxford Nanopore Technologies.
Only personalise run settings if you are an experienced user. Any deviations from the default settings have not been qualified and are done at the operators risk.
If using personalised volume and timer inputs:
- Select 'No' under 'Use standard volume and timers'.
Note: All categories will be autofilled with the recommended settings. These can be edited for personal requirements.
- Enter the desired reagent volumes for each sample in the 'End-repair' section I and II.
- Enter the desired length for each process in the 'End-repair' section of the protocol.
- Enter the desired reagent volumes for each sample in the 'Adapter ligation' section I and II.
- Enter the desired length for each process in the 'End-repair' section of the protocol.
Review the run variables and click 'Confirm':

Follow the on-screen directions to load the 150 µl filtered tips for the Multichannel Arm 384/96 onto the worktable:
- The required loading position for each box will flash to indicate where to place the labware.
- Please load full tip boxes only, partially full boxes of filtered tips for the Multichannel Arm 384/96 are currently not supported.
- Click 'Approve' after each addition of labware to proceed to the next box.
- After loading all of the required labware, close the front safety shield and click 'Next Page' to proceed.

Follow the on-screen directions to load the 50 µl filtered tips for the Multichannel Arm 384/96 onto the worktable:
- The required loading position for each box will flash to indicate where to place the labware.
- Please load full tip boxes only, partially full boxes of filtered tips for the Multichannel Arm 384/96 are currently not supported.
- Click 'Approve' after each addition of labware to proceed to the next box.
- After loading all of the required labware, close the front safety shield and click 'Next Page' to proceed.

Follow the on-screen directions to load the 1000 µl Flexible Channel Arm filtered tips onto the worktable:
- The required loading position for each box will flash to indicate where to place the labware.
- Partially full boxes are supported for the Flexible Channel Arm filtered tips. However, please ensure the tip box contains the minimum required number of tips as described in the equipment and consumables section of this protocol.
- Click 'Approve' after each addition of labware to proceed to the next box.

Follow the on-screen directions to load the 200 µl Flexible Channel Arm filtered tips onto the worktable:
- The required loading position for each box will flash to indicate where to place the labware.
- Partially full boxes are supported for the Flexible Channel Arm filtered tips. However, please ensure the tip box contains the minimum required number of tips as described in the equipment and consumables section of this protocol.
- Click 'Approve' after each addition of labware to proceed to the next box.

Follow the on-screen directions to load the 50 µl Flexible Channel Arm filtered tips onto the worktable:
- The required loading position for each box will flash to indicate where to place the labware.
- Partially full boxes are supported for the Flexible Channel Arm filtered tips. However, please ensure the tip box contains the minimum required number of tips as described in the equipment and consumables section of this protocol.
- Click 'Approve' after each addition of labware to proceed to the next box.
- After loading all of the required labware, click 'Next Page' to proceed.

Follow the on-screen directions to load the metal lid for ODTC on to the worktable:
- The required loading position will flash to indicate where to place the labware.
- Click 'Approve' after the addition of the metal lid for ODTC to proceed.
Loading the metal lid for ODTC:

Note: Take care to position the metal lid centrally in the recess. Incorrect positioning of the metal lid can lead to run error.
Follow the on-screen directions to load the Waste plate on to the worktable:
- The required loading position will flash to indicate where to place the labware.
- After loading the Waste plate, click 'Next Page'.
Loading the Waste Plate:

Follow the on-screen directions to load the reaction plates onto the worktable:
Incorrect positioning of the reaction plates will result in incorrect sample tracking throughout the run. Please ensure the reaction plates are placed on the worktable in the correct orientation:
- For the reaction plate(s) loaded in the 'Hotel': The lettered well markers (A-H) should be positioned towards the back of the worktable and the numbered well markers (1-12) should be positioned facing the right.
- For the reaction plate(s) loaded onto the worktable: Follow standard plate orientation, with the lettered well markers (A-H) positioned to the left and the numbered well markers (1-12) positioned to towards the back of the worktable.
Note: We recommend all plates are labelled before placing on the worktable to ensure correct plate tracking.
- The required loading position for each plate will flash on the on-screen display to indicate where to place the labware.
- Click 'Approve' after each addition of labware to proceed.
- After loading all of the required labware, click 'Next Page'.

Follow the on-screen directions to load the Bead plate, the Elution buffer plate and Ethanol plate on to the worktable:
- The required loading position will flash to indicate where to place the labware.
- Click 'Approve' after each addition of labware to proceed.
- After loading all of the required labware, click 'Next Page'.
Note: We recommend all plates are labelled before placing on the worktable to ensure correct plate tracking.
Loading the 'Bead plate':

Loading the 'Elution buffer plate':

Loading the 'Ethanol plate':

Close the front safety shield of the Tecan DreamPrep NGS while you prepare the reaction master mixes off-deck.

Prepare the End-prep (EP) master mix in a 2 ml Sarstedt tube with the following reagents according to the Tecan DreamPrep NGS user interface. Click 'OK' and 'Continue' to proceed.
We recommend master mixes are prepared fresh following the instructions below and loaded onto the workspace as soon as possible for optimal results.
- Ensure the master mix is homogenous by pipette mixing.
- Avoid the introduction of air bubbles while preparing the master mix.
- Avoid leaving droplets on the tube wall.
- Briefly spin down the End-prep (EP) master mix to ensure all the liquid is at the bottom of the Sarstedt tube.
Reagent volumes for all sample numbers:
| Reagent | Volume per sample | Volume X8 samples | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|---|---|
| NEBNext FFPE DNA Repair Buffer | 3.5 µl | 36.4 µl | 109.2 µl | 201.6 µl | 369.6 µl |
| NEBNext FFPE DNA Repair Mix | 2 µl | 20.8 µl | 62.4 µl | 115.2 µl | 211.2 µl |
| Ultra II End-prep reaction buffer | 3.5 µl | 36.4 µl | 109.2 µl | 201.6 µl | 369.6 µl |
| Ultra II End-prep enzyme mix | 3 µl | 31.2 µl | 93.6 µl | 172.8 µl | 316.8 µl |
| Total | 12 µl | 124.8 µl | 374.4 µl | 691.2 µl | 1267.2 µl |
Note: Reagent volumes will vary in accordance with the number of samples selected for processing (8-96). If processing a different sample input numbers, follow the instructions provided by the on-screen display for the correct reagent volumes. Volumes indicated in the table and the on-screen display will include the necessary dead volume excess.
Prepare the Adapter-ligation (AL) master mix directly into the 2 ml Sarstedt tube(s) with the following reagents according to the Tecan DreamPrep NGS user interface. Click 'OK' and 'Continue' to proceed.
We recommend master mixes are prepared fresh following the instructions below and loaded onto the workspace as soon as possible for optimal results.
- Ensure the master mix is homogenous by pipette mixing.
- Avoid the introduction of air bubbles while preparing the Adapter-ligation (AL) master mix.
- Avoid leaving droplets on the tube wall.
- Briefly spin down the Adapter-ligation (AL) master mix to ensure all the liquid is at the bottom of the Sarstedt tube(s).
Reagent volumes for preparation in each tube for all sample numbers:
| Reagent | Volume per sample | Volume X8 samples | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|---|---|
| Number of tubes to prepare | - | 1 | 1 | 2 | 4 |
| Ligation Buffer (LNB) | 25 µl | 260 µl | 780 µl | 660 µl | 660 µl |
| NEBNext Quick T4 DNA Ligase | 10 µl | 104 µl | 312 µl | 264 µl | 264 µl |
| Ligation Adapter (LA) | 5 µl | 52 µl | 156 µl | 132 µl | 132 µl |
| Total volume in each tube | - | 416 µl | 1,248 µl | 1,056 µl | 1,056 µl |
| Total volume prepared | - | 416 µl | 1,248 µl | 2,112 µl | 4,224 µl |
Note: Reagent volumes will vary in accordance with the number of samples selected for processing (8-96). If processing a different sample input numbers, follow the instructions provided by the on-screen display for the correct reagent volumes and split accordingly across the Sarstedt tube(s). Volumes indicated in the table and the on-screen display will include the necessary dead volume excess.
Load the DNA repair and end-prep master mix and Adapter ligation master mix prepared above in the 2 ml Sarstedt tubes into the required positions in the POGO tube holder by following the on-screen instructions.
- Ensure the master mixes are thoroughly mixed before loading.
- You will need to select each loading position using the Tecan's TouchTools touchscreen display.
- Follow the instructions on the display for each reagent, ensuring the fill volume for each tube is correct.
- Ensure the reagents have been added to all of the required positions before proceeding.
- Click 'Confirm' to proceed.
On-screen abbreviation glossary:
- End-prep master mix – EP
- Adapter ligation master mix – AL

Please ensure the liquid is evenly distributed across the troughs.

Gently tilt liquid in the trough backwards and forward a few times to generate a wet surface and allow the full volume to be evenly distributed.
Uneven distribution of the volume in the trough can be detrimental to the run.
Trough mounts and loading:
Trough mount sites:

Trough loading:
For the 100 ml troughs, dispense the required reagent volume directly into the trough and load on the the worktable. The 100 ml trough use will be reflected on the screen display with the following image:

For the 25 ml troughs, dispense the required reagent volume into the trough. You will then need to insert the 25 ml trough into a 100 ml trough to act as a holder or use a re-useable metal insert (for more information on additional equipment contact your Tecan representative). The modified 25 ml trough containing the reagent will then be loaded into the worktable. This input will be reflected on the screen display with the following image:

Depending on the wash buffer (LFB or SFB) used, the clean-up step after adapter ligation is designed to either enrich for DNA fragments of >3 kb, or purify all fragments equally.
- To enrich for DNA fragments of 3 kb or longer, use Long Fragment Buffer (LFB)
- To retain DNA fragments of all sizes, use Short Fragment Buffer (SFB)
Load the troughs with their relevant reagents into the worktable by following the on-screen instructions.
- Ensure all the reagents have been thoroughly mixed by vortexing before dispensing into the troughs.
- The required loading position will flash to indicate where to insert the trough.
- Click 'Approve' after each addition to proceed.
- After loading all of the troughs, click 'Next Page'.
Note: Follow the on-screen instructions for the reagent fill volume and the required trough to use.
- Site 2: Trough with fresh 80% ethanol.
- Site 3: Trough with AMPure XP Beads.
- Site 5: Trough with Long Fragment Buffer (LFB) or Short Fragment Buffer (SFB), depending on use.
- Site 6: Trough with Elution Buffer (EB).
- Site 7: Trough with nuclease-free water.
Close the front safety shield of the Tecan DreamPrep NGS while you prepare the sample plate off-deck.

Follow the on-screen directions to load the sample plate on to the worktable:
- Quantify your sample input using a Qubit fluorometer (or equivalent).
- Per sample, transfer 1 μg (or 100-200 fmol) of input DNA into a well of the input plate.
- Adjust the volume to 60 μl with nuclease-free water.
- Mix thoroughly by pipetting.
- Spin down briefly in a microfuge.
- The required loading position will flash to indicate where to place the labware.
- Click 'Approve' after the addition of labware to proceed.
- After loading all of the required labware, click 'Next Page'.

If removing the sample plate for off-deck storage, user interaction is required approximately 10 minutes after starting the run.

Close the front safety shield of the Tecan DreamPrep NGS before starting your run and click 'Continue'.

Click 'Confirm' and 'Continue' to start the automated library preparation.

Once the run has finished on the Tecan DreamPrep NGS remove your elution plate as soon as possible from the worktable to avoid evaporation.
We do not recommend running the liquid handling robot overnight as leaving the eluted library plate unsealed can lead to evaporation of the library product.
Please refer to the 'Timings' table found in the overview of the protocol for guidance.
Remove the plate containing the eluted libraries from the Tecan DreamPrep NGS deck.
Quantify 1 µl of each eluted sample from the output plate using a Qubit fluorometer plate reader off deck.
Seal the plate and store on ice until ready to prepare the library/libraries and load onto the flow cell.
We do not recommend running the liquid handling robot overnight as the plate must be sealed and stored on ice as soon as library preparation is finished.
Depending on your DNA library fragment size, prepare your final library in 12 µl of Elution Buffer (EB).
| Fragment library length | Flow cell loading amount |
|---|---|
| Very short (<1 kb) | 100 fmol |
| Short (1-10 kb) | 35–50 fmol |
| Long (>10 kb) | 300 ng |
Note: If the library yields are below the input recommendations, load the entire library.
If required, we recommend using a mass to mol calculator such as the NEB calculator.
We recommend loading the following amount of prepared library onto the R10.4.1 flow cell:
For high output, load 35-50 fmol of final library. This is to ensure high pore occupancy of >95% is reached. How to calculate pore occupancy can be found here.
Library storage recommendations
We recommend storing libraries at 4°C for short term storage or repeated use, for example, re-loading flow cells between washes. For single use and long term storage of more than 3 months, we recommend storing libraries at -80°C. For further information, please refer to the Library Stability Know-How document.
4. Priming and loading the MinION/GridION Flow Cell
Materials
- Flow Cell Flush (FCF)
- Flow Cell Tether (FCT)
- Library Solution (LIS)
- Library Beads (LIB)
- Sequencing Buffer (SB)
Consumables
- 1.5 ml Eppendorf DNA LoBind tubes
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Bovine Serum Albumin (BSA) (50 mg/ml) (e.g. Invitrogen™ UltraPure™ BSA 50 mg/ml, AM2616)
Equipment
- MinION or GridION device
- SpotON Flow Cell
- MinION/GridION Flow Cell Light Shield
- P1000 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
Please note, this kit is only compatible with R10.4.1 flow cells (FLO-MIN114).
Take the flow cell out of the fridge and leave it at room temperature for 20 minutes. This will improve visibility of the array during priming and sample loading.
Priming and loading a flow cell
We recommend all new users watch the 'Priming and loading your flow cell' video before your first run.
Using the Library Solution
For most sequencing experiments, use the Library Beads (LIB) for loading your library onto the flow cell. However, for viscous libraries it may be difficult to load with the beads and may be appropriate to load using the Library Solution (LIS).
Thaw the Sequencing Buffer (SB), Library Beads (LIB) or Library Solution (LIS, if using), Flow Cell Tether (FCT) and Flow Cell Flush (FCF) at room temperature before mixing by vortexing. Then spin down and store on ice.
For optimal sequencing performance and improved output on MinION R10.4.1 Flow Cells (FLO-MIN114), add Bovine Serum Albumin (BSA) to the flow cell priming mix at a final concentration of 0.2 mg/ml.
Note: We do not recommend using any other albumin type (e.g. recombinant human serum albumin).
Prepare the flow cell priming mix with BSA in a suitable vial for the number of flow cells to flush. Once combined, mix well by inverting the tube and pipette mix at room temperature.
| Reagent | Volume per flow cell |
|---|---|
| Flow Cell Tether (FCT) | 30 µl |
| Flow Cell Flush (FCF) | 1170 µl |
| Bovine Serum Albumin (BSA) at 50 mg/ml | 5 µl |
| Total | 1,205 µl |
Open the MinION or GridION device lid and slide the flow cell under the clip. Press down firmly on the priming port cover to ensure correct thermal and electrical contact.
Complete a flow cell check to assess the number of pores available before loading the library.
This step can be omitted if the flow cell has been checked previously.
See the flow cell check document for more information.
Slide the flow cell priming port cover clockwise to open the priming port.
Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µl, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the priming port
- Turn the wheel until the dial shows 220-230 µl, to draw back 20-30 µl, or until you can see a small volume of buffer entering the pipette tip
Note: Visually check that there is continuous buffer from the priming port across the sensor array.
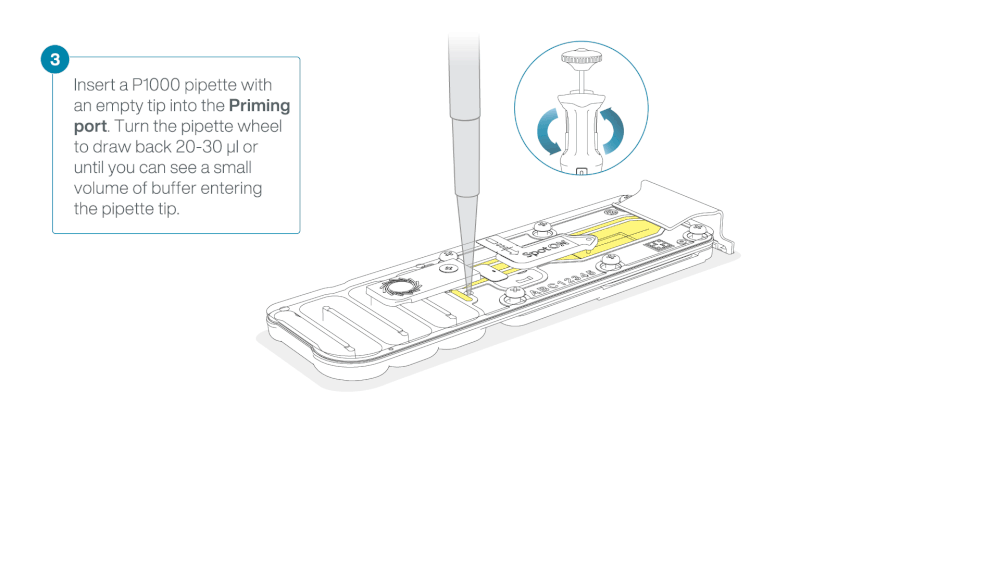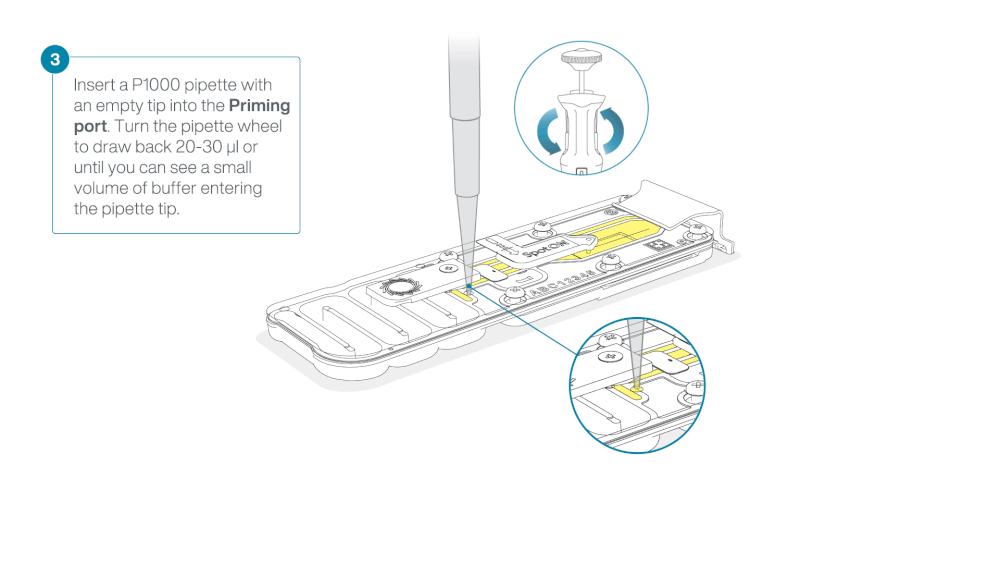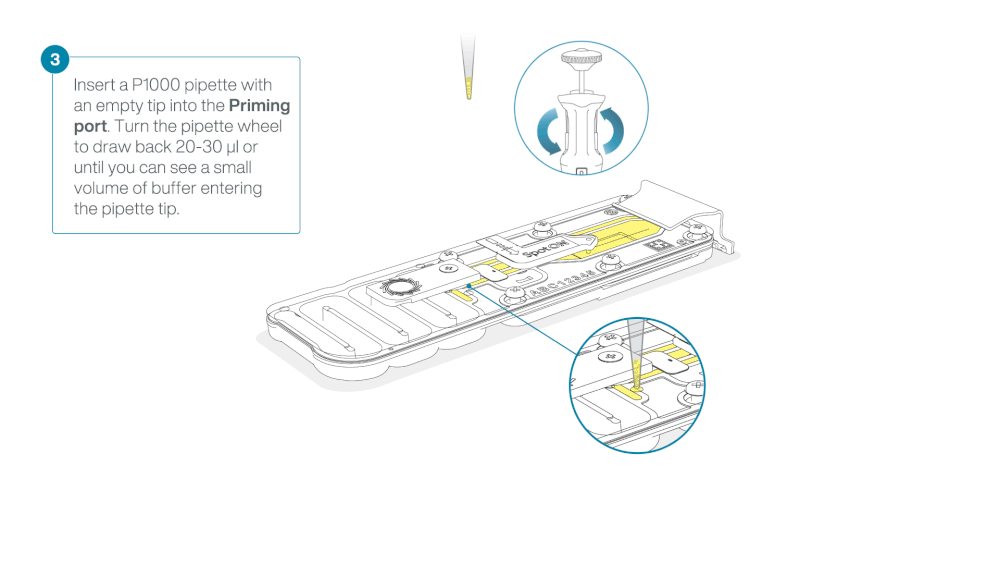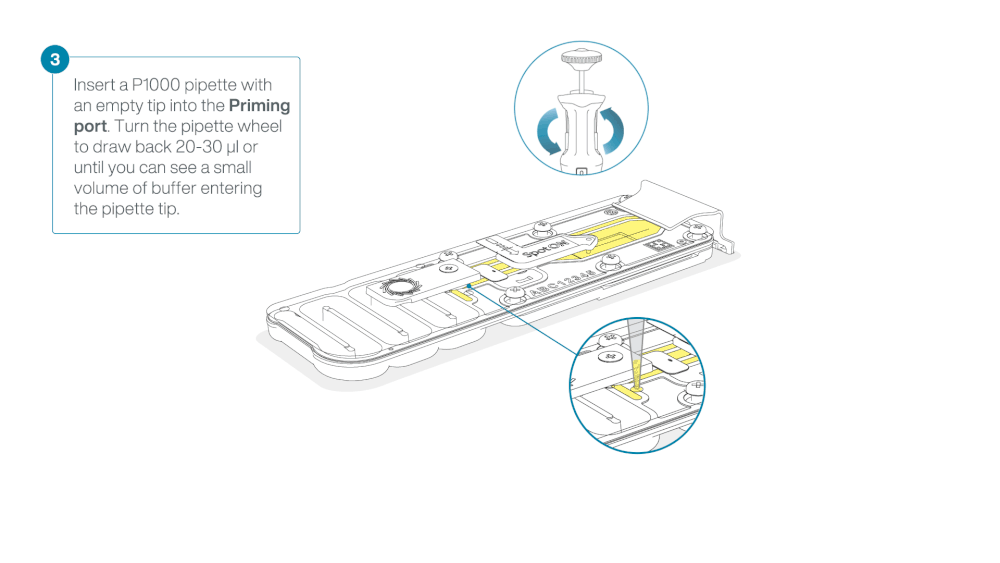
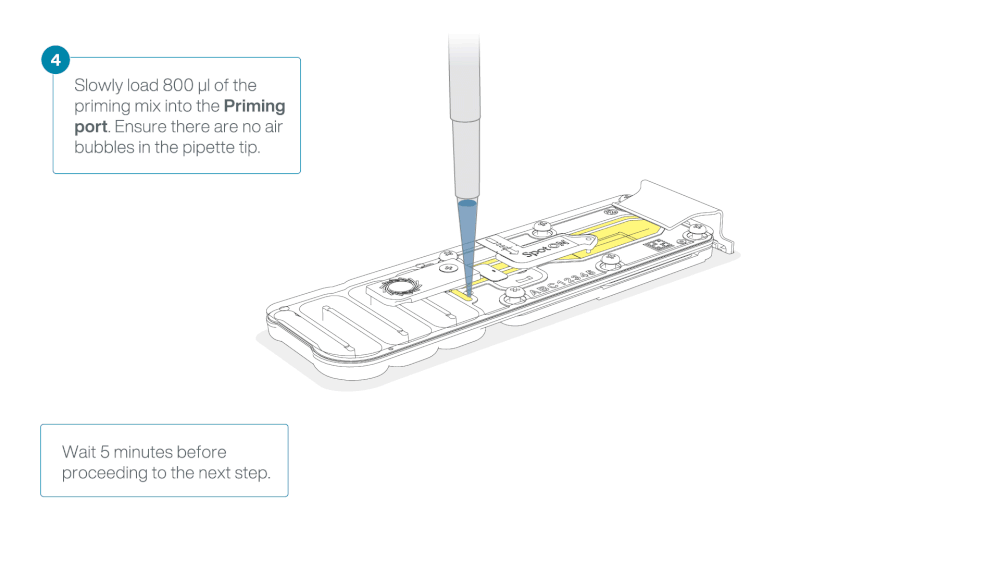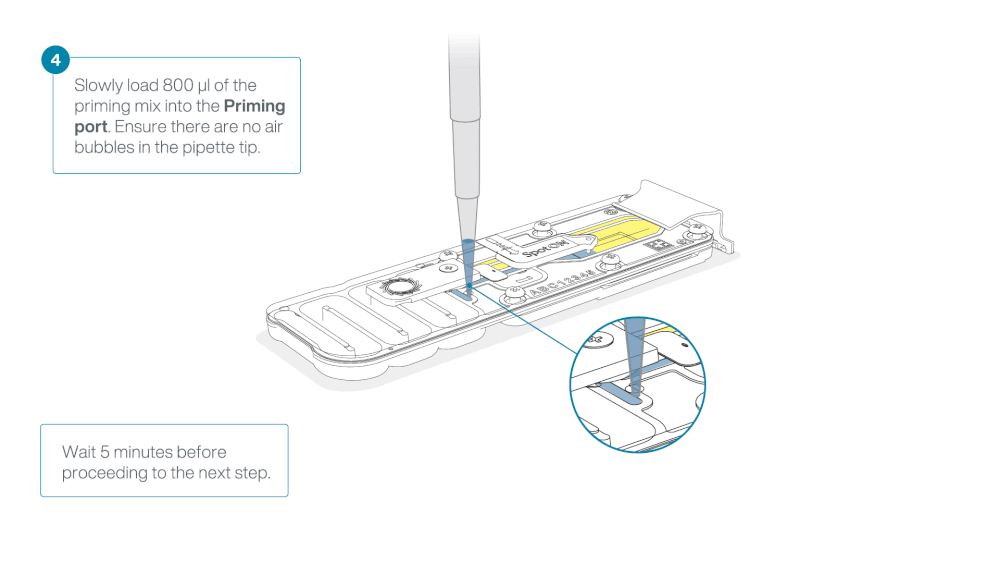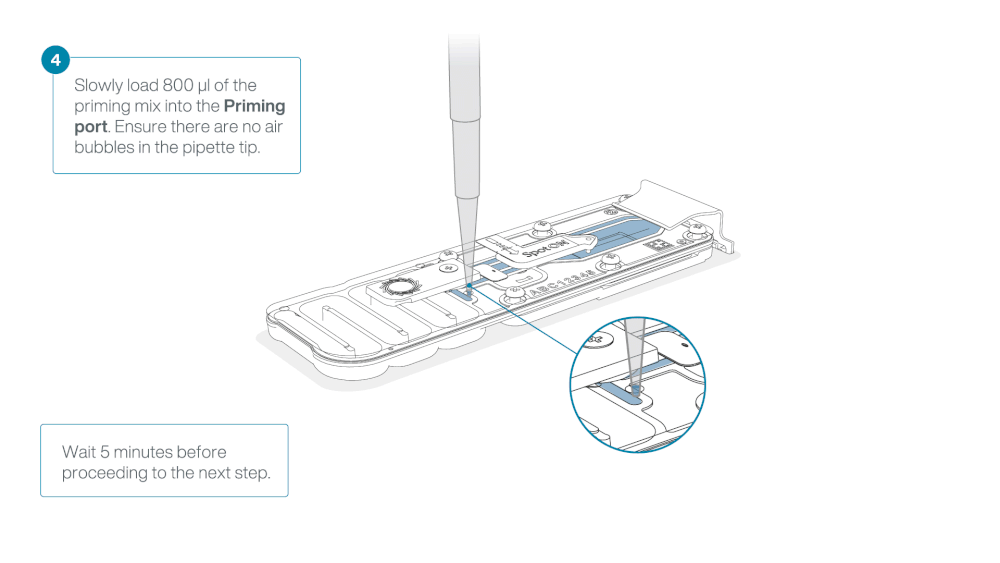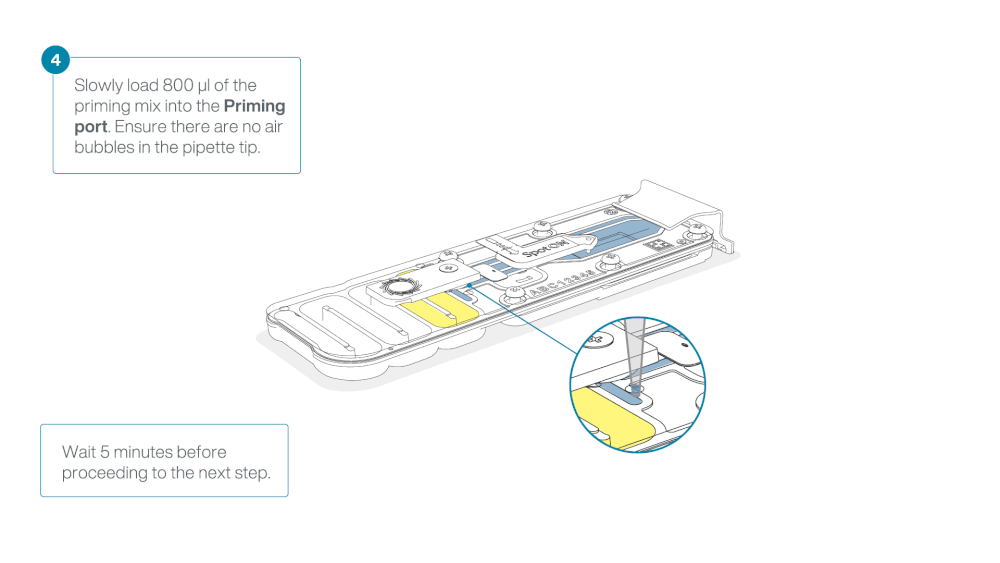
Load 800 µl of the priming mix into the flow cell via the priming port, avoiding the introduction of air bubbles. Wait for five minutes. During this time, prepare the library for loading by following the steps below.
Thoroughly mix the contents of the Library Beads (LIB) by pipetting.
The Library Beads (LIB) tube contains a suspension of beads. These beads settle very quickly. It is vital that they are mixed immediately before use.
We recommend using the Library Beads (LIB) for most sequencing experiments. However, the Library Solution (LIS) is available for more viscous libraries.
In a new 1.5 ml Eppendorf DNA LoBind tube, prepare the library for loading as follows:
| Reagent | Volume per flow cell |
|---|---|
| Sequencing Buffer (SB) | 37.5 µl |
| Library Beads (LIB) mixed immediately before use, or Library Solution (LIS), if using | 25.5 µl |
| DNA library | 12 µl |
| Total | 75 µl |
Complete the flow cell priming:
- Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
- Load 200 µl of the priming mix into the flow cell priming port (not the SpotON sample port), avoiding the introduction of air bubbles.

Mix the prepared library gently by pipetting up and down just prior to loading.
Add 75 μl of the prepared library to the flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.

Gently replace the SpotON sample port cover, making sure the bung enters the SpotON port and close the priming port.

For optimal sequencing output, install the light shield on your flow cell as soon as the library has been loaded.
We recommend leaving the light shield on the flow cell when library is loaded, including during any washing and reloading steps. The shield can be removed when the library has been removed from the flow cell.
Place the light shield onto the flow cell, as follows:
Carefully place the leading edge of the light shield against the clip. Note: Do not force the light shield underneath the clip.
Gently lower the light shield onto the flow cell. The light shield should sit around the SpotON cover, covering the entire top section of the flow cell.

The MinION Flow Cell Light Shield is not secured to the flow cell and careful handling is required after installation.
Close the device lid and set up a sequencing run on MinKNOW.
When a flow cell is inserted into the MinION Mk1D, the device lid will sit on top of the flow cell, leaving a small gap around the sides. This is normal and has no impact on the performance of the device.
Please refer to this FAQ regarding the device lid.

5. Unloading the Tecan DreamPrep NGS worktable
Consumables
- Low-lint scientific wipes (e.g. Kimberly-Clark™, 7552 or equivalent)
- Double distilled water (ddH2O) (e.g. ThermoFisher, 11983084)
- Freshly prepared ≥80% ethanol in nuclease-free or double distilled water, for cleaning
- ~10% Bleach (or equivalent): Thermo Scientific Alfa Aesar Sodium hypochlorite, 11-15% available chlorine, (e.g. ThermoFisher, 15429019)
Please ensure you have removed the plate containing your eluted sample libraries and stored it appropriately before unloading the rest of the worktable.
Empty the tip waste container.
Dispose of the used tips in an appropriate container.
Remove all the remaining plates from the worktable and hotel sites, and discard accordingly.
Remove the disposable 25 ml and 100 ml troughs from the trough mounting sites, and discard accordingly.
Note: Take care not to spill any residual liquid waste when removing from the worktable.
Remove the Sarstedt tubes from the POGO tube holder, and discard accordingly.
Clean the Bio-Rad™ Arched Auto-Sealing Lid:
- Wipe the lid with 10% bleach
- Thoroughly rinse the bleach off the lid using ≥80% ethanol or double distilled water (ddH2O) and lint-free wipes
- Allow the lid to air dry
Remove and/or restock the tip boxes on the worktable:
- Remove all Flexible Channel Arm (FCA) hanging tip trays from the FCA standard tip carriers, and discard accordingly.
- Remove all empty MultiChannel Arm (MCA) tip boxes from the worktable and discard accordingly.
- For partially used tip boxes, consider restacking the box with the appropriate tips for the next run.
Note: The MultiChannel Arm (MCA) tip boxes must be fully stacked. Failure to fully stack the MCA tip boxes can result in run error.
In cases where spillage has occurred during the automated library preparation, wipe the worktable surface using ≥80% ethanol.
Conclude all open dialogues on the TouchTools screen.
6. Data acquisition and basecalling
How to start sequencing
Once you have loaded your flow cell, the sequencing run can be started on MinKNOW, our sequencing software that controls the device, data acquisition and real-time basecalling. For more detailed information on setting up and using MinKNOW, please see the MinKNOW protocol.
MinKNOW can be used and set up to sequence in multiple ways:
- On a computer either directly or remotely connected to a sequencing device.
- Directly on a GridION or PromethION 24/48 sequencing device.
For more information on using MinKNOW on a sequencing device, please see the device user manuals:
To start a sequencing run on MinKNOW:
1. Navigate to the start page and click Start sequencing.
2. Fill in your experiment details, such as name and flow cell position and sample ID.
3. Select the Ligation Sequencing Kit XL V14 (SQK-LSK114) on the Kit page.
4. Configure the sequencing and output parameters for your sequencing run or keep to the default settings on the Run configuration tab.
Note: If basecalling was turned off when a sequencing run was set up, basecalling can be performed post-run on MinKNOW. For more information, please see the MinKNOW protocol.
5. Click Start to initiate the sequencing run.
Data analysis after sequencing
After sequencing has completed on MinKNOW, the flow cell can be reused or returned, as outlined in the Flow cell reuse and returns section.
After sequencing and basecalling, the data can be analysed. For further information about options for basecalling and post-basecalling analysis, please refer to the Data Analysis document.
In the Downstream analysis section, we outline further options for analysing your data.
7. Flow cell reuse and returns
Materials
- Flow Cell Wash Kit (EXP-WSH004)
After your sequencing experiment is complete, if you would like to reuse the flow cell, please follow the Flow Cell Wash Kit protocol and store the washed flow cell at +2°C to +8°C.
The Flow Cell Wash Kit protocol is available on the Nanopore Community.
We recommend you to wash the flow cell as soon as possible after you stop the run. However, if this is not possible, leave the flow cell on the device and wash it the next day.
Alternatively, follow the returns procedure to send the flow cell back to Oxford Nanopore.
Instructions for returning flow cells can be found here.
If you encounter issues or have questions about your sequencing experiment, please refer to the Troubleshooting Guide that can be found in the online version of this protocol.
8. Downstream analysis
Post-basecalling analysis
There are several options for further analysing your basecalled data:
EPI2ME workflows
For in-depth data analysis, Oxford Nanopore Technologies offers a range of bioinformatics tutorials and workflows available in EPI2ME, which are available in the EPI2ME section of the Community. The platform provides a vehicle where workflows deposited in GitHub by our Research and Applications teams can be showcased with descriptive texts, functional bioinformatics code and example data.
Research analysis tools
Oxford Nanopore Technologies' Research division has created a number of analysis tools, that are available in the Oxford Nanopore GitHub repository. The tools are aimed at advanced users, and contain instructions for how to install and run the software. They are provided as-is, with minimal support.
Community-developed analysis tools
If a data analysis method for your research question is not provided in any of the resources above, please refer to the resource centre and search for bioinformatics tools for your application. Numerous members of the Nanopore Community have developed their own tools and pipelines for analysing nanopore sequencing data, most of which are available on GitHub. Please be aware that these tools are not supported by Oxford Nanopore Technologies, and are not guaranteed to be compatible with the latest chemistry/software configuration.
9. Issues during automation of library preparation
Please contact your local Tecan Helpdesk and/or Nanopore FAS if you have any issues.
Your Tecan Helpdesk can be located via the following link: https://www.tecan.com/contact-us
10. Issues during the sequencing run
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Fewer pores at the start of sequencing than after Flow Cell Check
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | An air bubble was introduced into the nanopore array | After the Flow Cell Check it is essential to remove any air bubbles near the priming port before priming the flow cell. If not removed, the air bubble can travel to the nanopore array and irreversibly damage the nanopores that have been exposed to air. The best practice to prevent this from happening is demonstrated in videos for how to load a MinION Flow Cell and how to load a PromethION Flow Cell. |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | The flow cell is not correctly inserted into the device | Stop the sequencing run, remove the flow cell from the sequencing device and insert it again, checking that the flow cell is firmly seated in the device and that it has reached the target temperature. If applicable, try a different position on the device (GridION/PromethION). |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | Contaminations in the library damaged or blocked the pores | The pore count during the Flow Cell Check is performed using the QC DNA molecules present in the flow cell storage buffer. At the start of sequencing, the library itself is used to estimate the number of active pores. Because of this, variability of about 10% in the number of pores is expected. A significantly lower pore count reported at the start of sequencing can be due to contaminants in the library that have damaged the membranes or blocked the pores. Alternative DNA/RNA extraction or purification methods may be needed to improve the purity of the input material. The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
MinKNOW script failed
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Script failed" | Restart the computer and then restart MinKNOW. If the issue persists, please collect the MinKNOW log files and contact Technical Support. If you do not have another sequencing device available, we recommend storing the flow cell and the loaded library at 4°C and contact Technical Support for further storage guidance. |
Pore occupancy below 40%
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Pore occupancy <40% | Not enough library was loaded on the flow cell | Ensure you load the recommended amount of good quality library in the relevant library prep protocol onto your flow cell. Please quantify the library before loading and calculate mols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to pmol" |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and sequencing adapters did not ligate to the DNA | Make sure to use the NEBNext Quick Ligation Module (E6056) and Oxford Nanopore Technologies Ligation Buffer (LNB, provided in the sequencing kit) at the sequencing adapter ligation step, and use the correct amount of each reagent. A Lambda control library can be prepared to test the integrity of the third-party reagents. |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and ethanol was used instead of LFB or SFB at the wash step after sequencing adapter ligation | Ethanol can denature the motor protein on the sequencing adapters. Make sure the LFB or SFB buffer was used after ligation of sequencing adapters. |
| Pore occupancy close to 0 | No tether on the flow cell | Tethers are adding during flow cell priming (FLT/FCT tube). Make sure FLT/FCT was added to FB/FCF before priming. |
Shorter than expected read length
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Shorter than expected read length | Unwanted fragmentation of DNA sample | Read length reflects input DNA fragment length. Input DNA can be fragmented during extraction and library prep. 1. Please review the Extraction Methods in the Nanopore Community for best practice for extraction. 2. Visualise the input DNA fragment length distribution on an agarose gel before proceeding to the library prep. 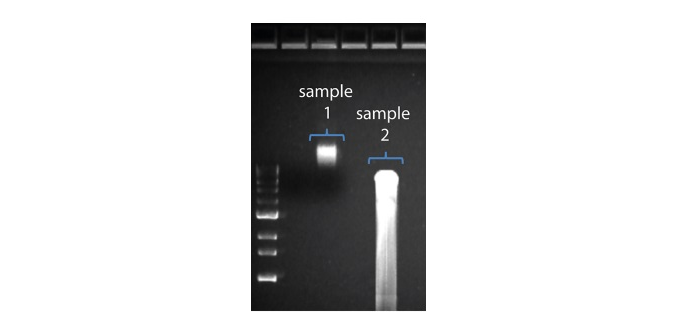 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented. In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.3. During library prep, avoid pipetting and vortexing when mixing reagents. Flicking or inverting the tube is sufficient. |
Large proportion of unavailable pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
Large proportion of unavailable pores (shown as blue in the channels panel and pore activity plot) 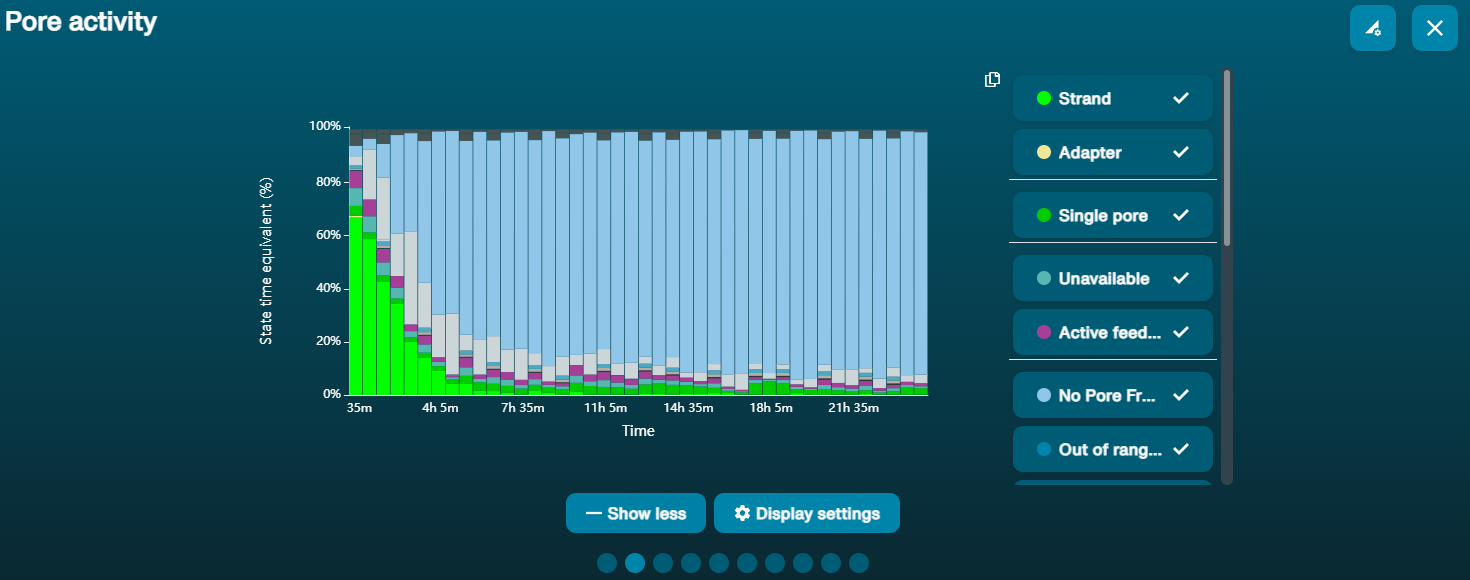 The pore activity plot above shows an increasing proportion of "unavailable" pores over time. The pore activity plot above shows an increasing proportion of "unavailable" pores over time. | Contaminants are present in the sample | Some contaminants can be cleared from the pores by the unblocking function built into MinKNOW. If this is successful, the pore status will change to "sequencing pore". If the portion of unavailable pores stays large or increases: 1. A nuclease flush using the Flow Cell Wash Kit (EXP-WSH004) can be performed, or 2. Run several cycles of PCR to try and dilute any contaminants that may be causing problems. |
Large proportion of inactive pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Large proportion of inactive/unavailable pores (shown as light blue in the channels panel and pore activity plot. Pores or membranes are irreversibly damaged) | Air bubbles have been introduced into the flow cell | Air bubbles introduced through flow cell priming and library loading can irreversibly damage the pores. Watch the how to load a MinION Flow Cell or how to load a PromethION Flow Cell videos for best practice. |
| Large proportion of inactive/unavailable pores | Certain compounds co-purified with DNA | Known compounds, include polysaccharides, typically associate with plant genomic DNA. 1. Please refer to the Plant leaf DNA extraction method. 2. Clean-up using the QIAGEN PowerClean Pro kit. 3. Perform a whole genome amplification with the original gDNA sample using the QIAGEN REPLI-g kit. |
| Large proportion of inactive/unavailable pores | Contaminants are present in the sample | The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
Reduction in sequencing speed and q-score later into the run
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Reduction in sequencing speed and q-score later into the run | For Kit 9 chemistry (e.g. SQK-LSK109), fast fuel consumption is typically seen when the flow cell is overloaded with library (please see the appropriate protocol for your DNA library to see the recommendation). | Add more fuel to the flow cell by following the instructions in the MinKNOW protocol. In future experiments, load lower amounts of library to the flow cell. |
Temperature fluctuation
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Temperature fluctuation | The flow cell has lost contact with the device | Check that there is a heat pad covering the metal plate on the back of the flow cell. Re-insert the flow cell and press it down to make sure the connector pins are firmly in contact with the device. If the problem persists, please contact Technical Services. |
Failed to reach target temperature
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Failed to reach target temperature" | The instrument was placed in a location that is colder than normal room temperature, or a location with poor ventilation (which leads to the flow cells overheating) | MinKNOW has a default timeframe for the flow cell to reach the target temperature. Once the timeframe is exceeded, an error message will appear and the sequencing experiment will continue. However, sequencing at an incorrect temperature may lead to a decrease in throughput and lower q-scores. Please adjust the location of the sequencing device to ensure that it is placed at room temperature with good ventilation, then re-start the process in MinKNOW. Please refer to this link for more information on MinION temperature control. |
11. This protocol is for Research Use Only. Not for use in diagnostic procedures.
Tecan Group Ltd. disclaimer
Tecan and DreamPrep are registered trademarks of Tecan Group Ltd., Männedorf, Switzerland. Flexible Channel Arm and MultiChannel Arm are trademarks of Tecan Group Ltd., Männedorf, Switzerland.
Tecan Group Ltd. makes every effort to include accurate and up-to-date information within this publication; however, it is possible that omissions or errors might have occurred. Tecan Group Ltd. cannot, therefore, make any representations or warranties, expressed or implied, as to the accuracy or completeness of the information provided in this publication. Changes in this publication can be made at any time without notice.
For technical details and detailed procedures of the specifications provided in this document please contact your Tecan representative.
In addition, this brochure may contain reference to applications and products which are not available in all markets.
For more information, please check with your Tecan local sales representative.







