Characterisation of drug resistance in Mycobacterium tuberculosis with targeted nanopore sequencing
- Published on: July 17 2023
Tuberculosis (TB), caused by the bacterium Mycobacterium tuberculosis, is the second highest cause of infectious disease-related deaths worldwide, second only to SARS-CoV-21, and caused 1.6 million deaths in 20212. TB can be classified by how resistant it is to drug treatment; an infection may be susceptible to all antibiotics, or it may be drug resistant (DR), multi-drug resistant (MDR) or extensively drug resistant (XDR). Application of the correct antibiotic therapy has consequences for disease outcome, as well as for the potential for antimicrobial resistance to arise.
Traditional TB resistance testing involves isolating the bacterium from respiratory samples and culturing it on agar plates in the presence of antibiotics. While growing cultures may only take days for some strains of bacteria, for M. tuberculosis this kind of phenotypic testing takes weeks, or sometimes months, due to its extremely slow growth rate. A multi-drug regimen is often used for treatment of TB before the pathogen has been characterised, but as the wrong treatment can result in treatment failure and contribute to further resistance, it is important to understand what kind of resistance phenotype an infection has as quickly as possible.
Resistance mutations can be determined using whole-genome sequencing (WGS) from respiratory samples, but one of the challenges of WGS is the need to wait weeks to culture enough bacteria for sequencing. A faster assay that could enable sequencing directly from respiratory samples, without the need for culture, was developed by Shannon Murphy and colleagues at the Wadsworth Center — New York State Department of Health, USA2. This culture-free, targeted sequencing method consisted of a multiplex PCR designed to amplify 13 target regions across nine antimicrobial resistance genes representing the first and second line treatments against TB. This was followed by library preparation, sequencing on a MinION Flow Cell using a MinION Mk1C or GridION in high-accuracy basecalling mode, and the use of a custom in-house analysis pipeline.
In her talk at the Nanopore Community Meeting 2022, Shannon explained that nanopore sequencing was selected for this application for its ability to generate long reads, enabling the use of fewer primer pairs and the ability to sequence entire amplicons, and added that ‘one of the most important reasons is the ability to have real-time monitoring and analysis of our data — this enables us to stop the run as soon we have sufficient data to report on, and we can ultimately get those results quicker’1.
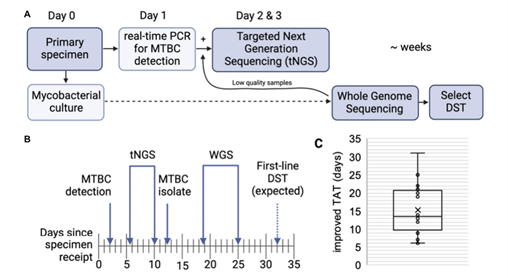
Figure 1: Turnaround times for MTBC molecular testing and sequencing. A) Overview of how specimens are processed. B) Timeline showing the average number of days required for MTBC DNA detection. C) Turnaround time improvements in days of direct sequencing compared with culture-derived WGS. Figure from Murphy et al.1 and available under the Creative Commons license (https://creativecommons.org/licenses/by/4.0/).
To investigate the future potential for the method to accurately characterise the resistance profile of M. tuberculosis infections, the team tested the specificity, sensitivity, reproducibility, and accuracy of the targeted assay. For sensitivity, the limit of detection was assessed using a spiked sputum sample; the targeted assay successfully identified all 13 gene targets down to 0.108 CFU per reaction. For reproducibility, results were found to be concordant between three replicates of three positive samples, sequenced on the same and different days. Specificity was checked against Mycobacterium fortuitum, M. abscessus, Klebsiella pneumoniae, Streptococcus pneumoniae, and Haemophilus influenzae and demonstrated that there was no cross-reactivity with these species in the assay.
Finally, accuracy was tested on 72 respiratory research samples; the resistance mutations identified were found to be 100% concordant with WGS. In addition, the targeted assay may provide the further benefit of identifying unknown novel mutations, particularly where samples contain both M. tuberculosis and M. avium, which can cause a failure to obtain a pure culture for WGS.
'... this assay was found to be accurate and generated susceptibility profiles comparable to those currently obtained with our existing WGS assay, which can only be performed on cultured isolates'
Murphy, S. et al.2
To further leverage the benefits of this method and gain further insights into the epidemiology of TB, the team designed a SNP-based lineage prediction tool from the gene targets of the assay. The algorithm was applied to the 72 research samples used in the study, to identify to which of seven main TB phylogenetic lineages each sample belonged; 98.2% of lineages were correctly identified using the targeted sequencing panel.
Furthermore, the assay satisfied one of the most important criteria for its development — on average, the targeted assay demonstrated a 15-day reduction in turnaround times versus WGS, with one XDR sample being classified in five days, versus an additional three weeks for the culture-based WGS results to be returned from the same sample. The authors concluded that 'this particular tNGS [targeted next generation sequencing] assay showed more than a two-week improvement in turnaround time compared to culture and WGS workflows at a similar cost’ and that ’this method also offers additional utility for cultures that are low quality for WGS analysis due to mixed organisms or low MTBC DNA concentration’.
Murphy, S. Profiling drug-resistant Mycobacterium tuberculosis with targeted nanopore sequencing. Presentation. Available at: https://nanoporetech.com/resource-centre/video/ncm22/profiling-drug-resistant-mycobacterium-tuberculosis-with-targeted-nanopore-sequencing (2022) [Accessed: 06 July 2023].
Murphy, S.G. et al. Direct detection of drug-resistant Mycobacterium tuberculosis using targeted next generation sequencing. Front. Public Health 11:1206056 DOI: https://doi.org/10.3389/fpubh.2023.1206056.







