Ending the diagnostic odyssey starts here
With an estimated 300 to 400 million people worldwide living with a rare disease1,2, the word ‘rare’ can feel like a contradiction.
Nearly 70% of rare diseases begin in childhood1. Years of growth and discovery are often replaced with repeated clinic visits, referrals to multiple specialists, and a long series of tests. It’s a journey families know all too well — the ‘diagnostic odyssey'.
Despite around 80% of rare diseases having a genetic cause1, fewer than 50%3-5 of individuals receive a confirmed genetic diagnosis. Devastatingly, 30% of children with a rare disorder die before their fifth birthday1.
For researchers and clinical genomics teams working in rare disease, the need is clear: find answers for more families, and find them sooner.
Writing in RARE Revolution Magazine, Elissa Levin, Oxford Nanopore, explained that while ‘not all genetic tests are created equal’, new technologies ‘could mean the difference between a family continuing a frustrating search and finally receiving clarity’6.
In this blog, we explore how Oxford Nanopore sequencing could play a pivotal role in shaping the future of rare disease genomics.
Longer reads. Richer insights. New possibilities.
Over the past decade, short-read sequencing, whether targeted panels, exomes, or whole genomes, has become embedded in rare disease research and clinical genomics workflows, and diagnostic yield has improved across many disorders.
Yet inherent limitations remain. Short-read technology struggles in repetitive or structurally complex regions of the genome. Structural variants, repeat expansions, phasing, and epigenetic modifications remain difficult to resolve.
An increasing number of studies now demonstrate that Oxford Nanopore technology holds the potential to address these limitations.
Because our technology can generate reads of any length — including ultra-long reads spanning megabases — you can access regions of the genome that are difficult to interrogate with short fragments, making it easier to resolve clinically relevant structural variants7.
The ability to detect methylation directly in the same sequencing run, and to generate results rapidly, means our technology can provide a faster and more comprehensive view of genomic variation — in one go.
It’s this broader perspective that has led clinician-researcher Danny Miller to state that ‘long-read sequencing will fundamentally change clinical genetic testing within five years ... even if the cost of generating other types of data falls to $0’8.
It’s a bold view. But the research momentum is building.
Looking beyond the limits of short reads
Here, we highlight a selection of recently published studies demonstrating how Oxford Nanopore sequencing is being applied in rare disease research. While each study addresses different disorders and variant types, they share a common theme: resolving questions that were previously unanswered.
1. Advancing long-read nanopore genome assembly and accurate variant calling for rare disease detection. AJHG (2025)
Negi et al. at the University of California, Santa Cruz, USA, investigated whether nanopore reads could accurately resolve disease-associated variants that were inaccessible to short-read sequencing. Each sample was sequenced on a single PromethION Flow Cell (~36x depth).
Our technology genetically resolved 26% (11/42) of cases, compared to 19% (8/42) with short-read sequencing. The authors emphasised the ‘potential to enhance diagnostic yield for rare monogenic diseases’, suggesting future utility in clinical genomics workflows.
Take-home: accessing previously inaccessible regions of the genome with Oxford Nanopore technology may help bridge the diagnostic gap in rare disease research.
2. Singleton rapid long-read genome sequencing as first tier genetic test for critically ill children with suspected genetic diseases. EJHG (2025).
This proof-of-concept study explored the use of nanopore sequencing as a potential first-tier genetic test for critically ill children with suspected genetic disease in intensive care.
After exome sequencing failed to provide a molecular diagnosis for two individuals, Kamolvisit et al. applied nanopore sequencing to all 18 children in their cohort. Causative pathogenic variants were identified in 11/18 individuals. Our technology also revealed additional genomic information, leading to reclassification of single nucleotide variants (SNVs) in three cases.
Take-home: with further validation and development, Oxford Nanopore sequencing shows potential as a first-tier diagnostic approach in time-sensitive settings.
3. Nanopore long-read sequencing as a first-tier diagnostic test to detect repeat expansions in neurological disorders. Int. J. Mol. Sci. (2025)
To establish a workflow for analysing short tandem repeats (STRs) in neurological disorders, de Boer et al. designed a custom nanopore sequencing panel targeting nine genes associated with spinocerebellar ataxia and fragile X syndrome.
Applying their 'raw data to report’ nanopore workflow to samples from 12 affected individuals, they achieved concordance with current diagnostic testing for 22/23 STRs. Importantly, the same assay enabled detection of SNVs, indels, and methylation patterns — in one go.
Take-home: Oxford Nanopore sequencing demonstrates comparable performance to existing methods for STR detection, but with additional variant types and methylation information in a single assay, it offers future potential for more integrated testing strategies.
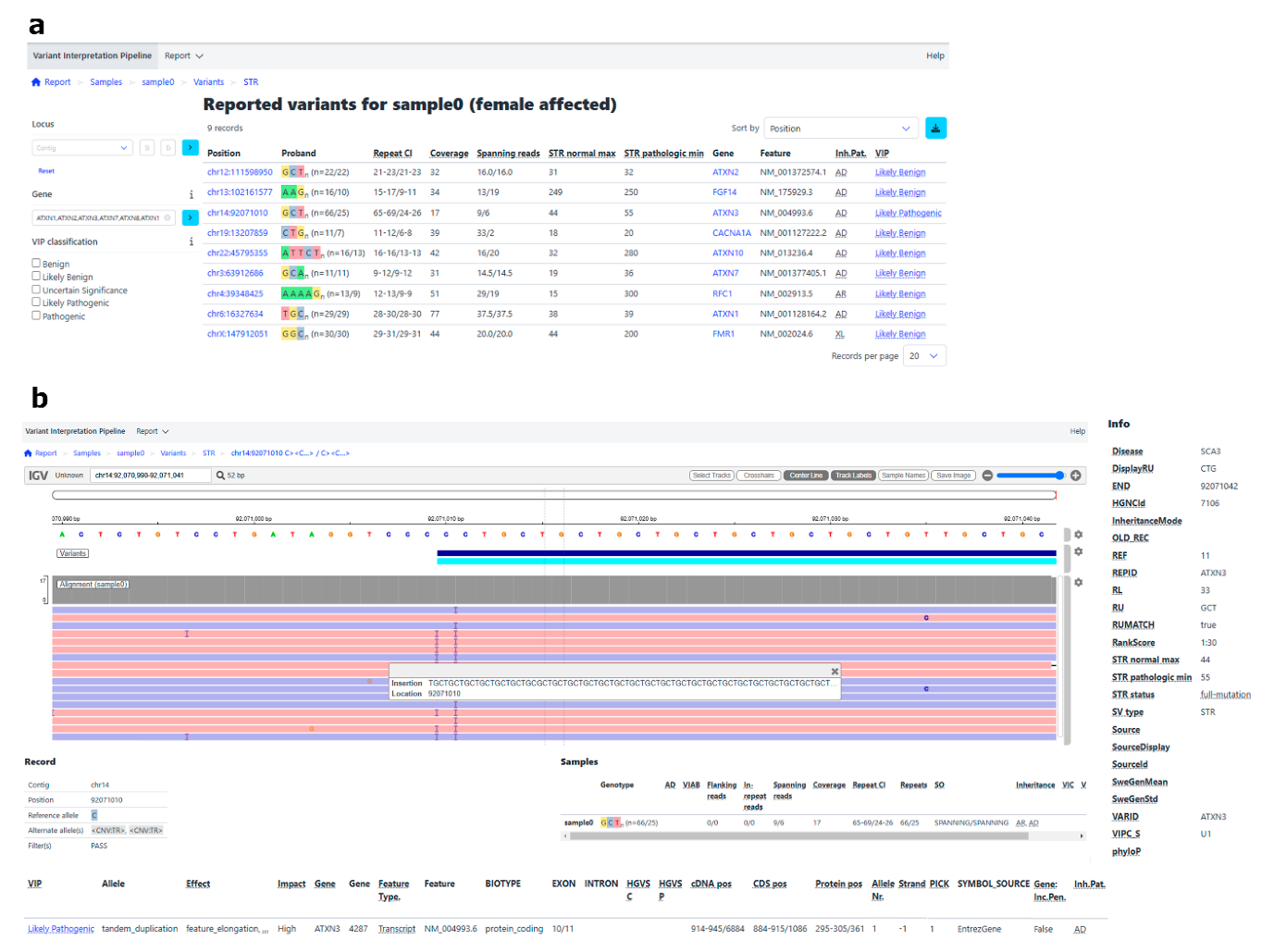
Figure: Variant interpretation report (VIP) which could be used in a clinical setting. The report shows results for targeted STRs (a), including the interpretation (far right column) and detailed results for a specific variant (b) — here being the ATXN3 repeat. Figure taken from de Boer et al. 2025 under Creative Commons Attribution License CC BY 4.0. The pre-print on MOLGENIS VIP provides further details on the pipeline.
4. Long-read sequencing enhances pathogenic and novel variation discovery in patients with rare diseases. Nat. Comm. (2025)
Sinha et al. applied whole-genome nanopore sequencing to a research cohort of undiagnosed individuals who had previously tested negative using short-read sequencing.
Among 51 individuals, additional potential diagnoses were uncovered in nearly 10% of cases (5/51). The team also identified a novel methylation signature relevant to spinal muscular atrophy, which could be incorporated into genetic testing in the future.
Take-home: for individuals with prior negative short-read results, Oxford Nanopore sequencing could reveal richer insights and increase diagnostic yield in the future.
Read senior author Ahmed Abou Tayoun’s story on developing a ‘unified platform for clinical genetic testing’.
5. Targeted long-read sequencing as a single assay improves the diagnosis of spastic-ataxia disorders. Ann. Clin. Transl. Neurol. (2025)
In genetically unresolved cases of spastic-ataxia, Rudaks et al. applied targeted Oxford Nanopore sequencing.
Causative variants were identified in 14/34 (41%) samples, including 9/23 (39%) cases that had remained undiagnosed after short-read sequencing. The authors suggest that targeted nanopore sequencing could ‘streamline the testing pathway [for spastic-ataxia] by capturing all known genetic causes in a single assay’.
Take-home: a consolidated Oxford Nanopore approach may simplify complex testing pathways in rare neurological disorders.
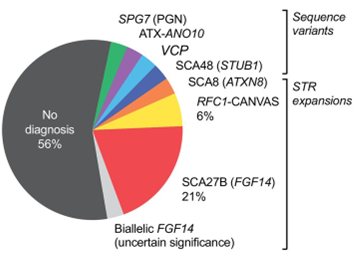
Figure: Overview of genetic findings for the research cohort of 34 individuals with spastic-ataxia spectrum disorders, including those with prior negative-genetic testing, and those who were testing naive. Figure taken from Rudaks et al. 2025 under Creative Commons License CC-BY 4.0.
6. The implementation of genome sequencing in rare genetic diseases diagnosis: a pilot study from the Hong Kong Genome Project The Lancet: Regional Health (2025)
In this prospective pilot study from the Hong Kong Genome Project, Lam et al. assessed the diagnostic yield of short-read sequencing followed by Oxford Nanopore sequencing.
Short-read sequencing identified causative variants in 24% (125/520) of cases. Our technology was subsequently applied to 21 samples with unresolved findings, identifying causative variation in all 21 cases.
Take-home: although currently a pilot study, Oxford Nanopore data could expand the proportion of genetically resolved rare disease cases in the future, with potentially great impact at population scale.
Building momentum for future rare disease diagnoses
Finally, we look to the BEACON project — the first large, independent, investigator-led clinical research initiative applying nanopore sequencing at scale in rare diseases.
The project includes 1,000 individuals across diverse rare disease categories, all previously unresolved after short-read sequencing. Among the first 141 cases analysed, 34 (24%) had their underlying genetic basis resolved.
Presenting the data at the Annual Clinical Genetics Meeting (ACMG) 2025, Wendy Chung of Boston Children’s Hospital reflected: ‘There’s no way in the world I would have picked this up from what we did before’.
The diagnostic odyssey has defined rare disease for decades. But it doesn’t have to define the future; Levin believes that ‘the journey from odyssey to answers is within reach’.
Oxford Nanopore technology is opening doors that once felt closed. Who knows, perhaps the next breakthrough is in your data.
Curious to learn more? Check out our 24-hour whole-genome sequencing workflow for rare disease research.
Oxford Nanopore Technologies products are not intended for use for health assessment or to diagnose, treat, mitigate, cure, or prevent any disease or condition.
FURTHER READING
Del Gobbo, G.F. and Boycott, K.M. The additional diagnostic yield of long-read sequencing in undiagnosed rare diseases. Genome Res. (2025). DOI: https://doi.org/10.1101/gr.279970.124
Damaraju, N., Miller, A.L., and Miller, D.E. Long-read DNA and RNA sequencing to streamline clinical genetic testing and reduce barriers to comprehensive genetic testing. J. Appl. Lab. Med. 9(1):138–150 (2024). DOI: https://doi.org/10.1093/jalm/jfad107
Paschal, C.R. et al. Concordance of whole-genome long-read sequencing with standard clinical testing for Prader-Willi and Angelman syndromes. J. Mol. Diagn. 27(3):166-176 (2024). DOI: https://doi.org/10.1016/j.jmoldx.2024.12.003
REFERENCES
The landscape for rare diseases in 2024. The Lancet Global Health 12(3):e341 (2024). DOI: https://doi.org/10.1016/S2214-109X(24)00056-1
Khoury, M.J. and Yu, W. Introducing the rare disease genomics and precision health knowledge base. https://blogs-origin.cdc.gov/genomics/2019/04/04/introducing-the-rare-diseases/ (2019) [Accessed 2 April 2025]
Alfares, A. et al. Whole-genome sequencing offers additional but limited clinical utility compared with reanalysis of whole-exome sequencing. Genet. Med. 20(11):1328–1333 (2018). DOI: https://doi.org/10.1038/gim.2018.41
Kingsmore, S.F. et al. A randomised, controlled trial of the analytic and diagnostic performance of singleton and trio, rapid genome and exome sequencing in ill infants. AJHG 105(4):719–733 (2019). DOI: https://doi.org/10.1016/j.ajhg.2019.08.009
Negi, S. et al. Advancing long-read nanopore genome assembly and accurate variant calling for rare disease detection. AJHG 112(2):428–449 (2025). DOI: https://doi.org/10.1016/j.ajhg.2025.01.002
Levin, E. From odyssey to answers. RARE Revolution Magazine 34:28–31 (2025). https://editions.rarerevolutionmagazine.com/html5/reader/production/default.aspx?pubname=&edid=5fddfc62-9dc6-47bf-aa6a-922d41bce312&pnum=29 [Accessed 17 February 2026]
Pei, Y. et al. A comparison of structural variant calling from short-read and nanopore-based whole-genome sequencing using optical genome mapping as a benchmark. Genes 15(7): 925 (2024). DOI: https://doi.org/10.3390/genes15070925
Miller, D. Long-read sequencing as the future of clinical genetic testing. Presentation. Available at: https://www.youtube.com/watch?v=9zaULIooEyI&ab_channel=OxfordNanoporeTechnologies (2024) [Accessed 17 February 2026]






