Data analysis (DATD_5000_v1_revZ_08Apr2026)
TechnicalDocument
V DATD_5000_v1_revZ_08Apr2026
FOR RESEARCH USE ONLY
1. Basecalling overview
Introduction to basecalling
Basecalling is the process of converting the electrical signals generated by a DNA or RNA strand passing through the nanopore into the corresponding base sequence of the strand. The general data flow in a nanopore sequencing experiment is shown below.
Raw data – a direct measurement of the changes in ionic current as a DNA/RNA strand passes through the pore, which are recorded by the MinKNOW software. MinKNOW also processes the signal into "reads", each read corresponding to a single strand of DNA/RNA. These reads are written out as POD5 files, a custom Oxford Nanopore file type.
Basecalling – the basecalling algorithm uses signal processing techniques based on machine learning to transform the raw signal of the reads into basecalls. The software writes out the results of these analyses into BAM files (unaligned, or containing modified base information and/or alignment information). Additionally, FASTQ files are also produced.
Oxford Nanopore provides several devices to allow users to carry out basecalling in real-time, as well as executables for users' local infrastructure. You can carry out basecalling live during the experiment, as post-processing following the completion of an experiment, or a combination of these.
Basecalling with neural networks
Oxford Nanopore basecallers convert raw signal data to basecalls using algorithms based on neural networks (a machine learning model).
These neural networks are loosely modelled on biological neural networks within the human brain. The network contains nodes (similar to neurons in the brain) arranged in layers, which carry out computations. Neural networks receive and process data, but crucially, they have been trained to have exceptional performance for particular signal processing tasks. They have been used successfully for diverse applications like pattern recognition (such as handwritten characters, speech recognition), or predicting trends over time.
Learn more about how basecalling works.
Basecalling model: 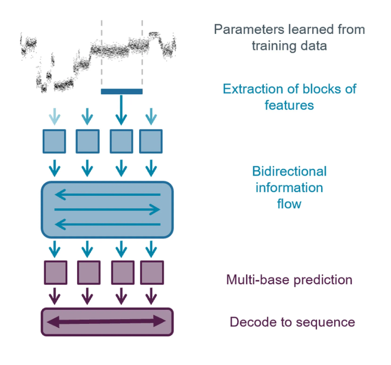
The neural networks used in Oxford Nanopore basecallers have been trained on a range of example DNA sequences (described in more detail in the Basecaller training section of Basecalling algorithms. The network learns how to translate the series of measurements into a sequence.
Oxford Nanopore basecallers
| Basecaller | Algorithm | Information |
|---|---|---|
| MinKNOW basecaller | Production basecaller on the device software. This is identical to the algorithm used by our stand-alone basecaller, but may be a version behind. | Available as a download (further details are in the section Downloading and installing MinKNOW in the MinKNOW protocol). Select the basecalling option when starting the sequencing experiment, and MinKNOW will display the experimental progress via the user interface. A Dorado-powered basecall server installed with MinKNOW is also available as a package for advanced users. Dorado is available as a download, and is also included in MinKNOW installations. |
| Dorado basecaller | Dorado is the production basecaller that is also available in MinKNOW. | Available as a download. You can run the executable version of the software on the host computer via the command line. |
2. Basecalling algorithms
Fast, High accuracy and Super accuracy models and compatibilities
The MinKNOW basecallers offer three different basecalling models: a Fast model, a High accuracy (HAC) model, and a Super accurate (SUP) model.
You will need to optimise accuracy according to your requirements by selecting the most suitable basecalling model.
Fast basecalling: fastest, least computationally intense. Compatible with real-time basecalling on all nanopore devices with compute. Recommended for quick, real-time insights on sequencing data when compute resources are limited.
High accuracy basecalling (HAC): highly accurate, intermediate speed and computational requirement. Compatible with real-time basecalling on GridION and PromethION devices with compute. Recommended for high-throughput projects focusing on variant analysis.
Super accurate basecalling (SUP): the most accurate and computationally intense. Recommended for de novo assembly projects and low-frequency variant analysis (e.g. somatic variation and single-cell applications).
The HAC model provides a higher raw read accuracy than the Fast model and is more computationally-intensive. The SUP model has an even higher raw read accuracy, and is even more intensive than the HAC model.
For more information about basecalling accuracy, see the Oxford Nanopore sequencing accuracy page.
A comparison of the speed of the models is provided for each device in the Tables below:
PromethION 24 A-series
| Number of keep-up flow cells for each task | Fast model | HAC model | SUP model |
|---|---|---|---|
| Basecalling only | 24 | 24 | 8 |
| Native barcoding 96 demultiplexing | 24 | 24 | 8 |
| Alignment | 24 | 24 | 8 |
| Modified bases 5mC + 5hmC (CpG context) | N/A | 24 | 8 |
| Modified bases 5mC + 5hmC (CpG context) with alignment | N/A | 22 | N/A |
PromethION 2 Integrated
| Number of keep-up flow cells for each task | Fast model | HAC model | SUP model |
|---|---|---|---|
| Basecalling only | 2 | 2 | 1.9 |
| Native barcoding 96 demultiplexing | 2 | 2 | 1.9 |
| Alignment | 2 | 2 | 1.9 |
| Modified bases 5mC + 5hmC (CpG context) | N/A | 2 | 1.9 |
| Modified bases 5mC + 5hmC (CpG context) with alignment | N/A | 2 | N/A |
PromethION 2 Solo connected to a GridION
| Number of keep-up flow cells for each task | Fast model | HAC model | SUP model |
|---|---|---|---|
| Basecalling only | 2 | 2 | 0.8 |
| Native barcoding 96 demultiplexing | 2 | 2 | 0.8 |
| Alignment | 2 | 2 | 0.8 |
| Modified bases 5mC + 5hmC (CpG context) | N/A | 2 | 0.8 |
| Modified bases 5mC + 5hmC (CpG context) with alignment | N/A | 2 | N/A |
GridION
| Number of keep-up flow cells for each task | Fast model | HAC model | SUP model |
|---|---|---|---|
| Basecalling only | 4.8 | 4.8 | 1.9 |
| Native barcoding 96 demultiplexing | 5 | 4.7 | 1.8 |
| Alignment | 4.8 | 4.6 | 1.8 |
| Modified bases 5mC + 5hmC (CpG context) | N/A | 4.7 | 1.6 |
| Modified bases 5mC + 5hmC (CpG context) with alignment | N/A | 4.7 | N/A |
Standalone MinKNOW on Apple M3 (MinION Flow Cells)
| Number of keep-up flow cells for each task | Fast model | HAC model | SUP model |
|---|---|---|---|
| Basecalling only | 1 | 1 | 0.2 |
| Native barcoding 96 demultiplexing | 1 | 1 | 0.2 |
| Alignment | 1 | 1 | 0.2 |
| Modified bases 5mC + 5hmC (CpG context) | N/A | 1 | 0.2 |
| Modified bases 5mC + 5hmC (CpG context) with alignment | N/A | 1 | N/A |
Standalone MinKNOW on NVIDIA GeForce RTX 5090 (MinION Flow Cells)
| Number of keep-up flow cells for each task | Fast model | HAC model | SUP model |
|---|---|---|---|
| Basecalling only | 1 | 1 | 1 |
| Native barcoding 96 demultiplexing | 1 | 1 | 1 |
| Alignment | 1 | 1 | 1 |
| Modified bases 5mC + 5hmC (CpG context) | N/A | 1 | 1 |
| Modified bases 5mC + 5hmC (CpG context) with alignment | N/A | 1 | N/A |
Standalone MinKNOW on NVIDIA GeForce RTX 5090 (PromethION Flow Cells)
| Number of keep-up flow cells for each task | Fast model | HAC model | SUP model |
|---|---|---|---|
| Basecalling only | 2 | 2 | 0.8 |
| Native barcoding 96 demultiplexing | 2 | 2 | 0.8 |
| Alignment | 2 | 2 | 0.8 |
| Modified bases 5mC + 5hmC (CpG context) | N/A | 2 | 0.8 |
| Modified bases 5mC + 5hmC (CpG context) with alignment | N/A | 2 | N/A |
The number of keep-up flow cells assumes 30 Gb of flow cell output in 72 hours for MinION and GridION, and 100 Gb output in 72 hours for PromethION. These benchmarks were carried out on 30 kb human DNA.
MinKNOW basecalling: keep-up vs catch-up
Basecalling with the Fast basecalling model can keep up with the speed of data acquisition on most of Oxford Nanopore devices. When using the more computationally-intensive models, basecalling continues after the sequencing experiment has run to completion; any reads that have not been basecalled during the experiment will be queued and processed afterwards. This is known as “Catch-up mode”.
You therefore have two options: either to allow MinKNOW to continue in catch-up mode, or to stop the analysis and basecall the remaining reads at a later time, e.g. using standalone Dorado.

Calling modified bases
Oxford Nanopore’s technology allows for direct, real-time sequencing and detection of modified bases for both DNA and RNA (e.g. 5mC, 5hmC, 6mA, 4mC for DNA, m6A, and pseudoU for RNA) without additional experiments or preparation. This requires the use of a designated basecalling model that is trained to identify base modifications. The simplest way to access these models is via MinKNOW on the device, where modified bases can be selected under the basecalling options for a sequencing run or via the standalone Dorado basecaller from GitHub.
For best practices and benchmarking for modified bases, please refer to DNA modified bases and RNA modified bases. Please refer to the base modification accuracy section on the sequencing accuracy page for more information.
Several advanced options are also available for calling and analysing modified bases. Use Modkit (also available on GitHub) for post-processing base modifications after basecalling. Modkit creates summary counts of modified and unmodified bases in an extended bedMethyl format. The bedMethyl files tabulate the counts of base modifications from every sequencing read over each reference genomic position.
Basecaller, consensus and variant caller model training
When developing basecalling, consensus, and variant-calling models using machine learning, Oxford Nanopore uses data from sequencing experiments. This data can be synthetic or derived from genomic sources. Model development is broken down into two broad categories: training (creating a model) and validation (showing that it works).
The data includes both PCR-amplified reads and native reads that can contain base modifications. A portion of the reads and/or genomic locations are reserved for validating the model and not included in the training dataset.
Once trained, the quality of the model is validated using reads covering genomic regions that were not included in the training dataset. Validation assesses the following parameters:
- Alignment accuracy
- Percentage of strands that align to the reference
- Identifying strand edges and barcodes
- Specific test cases such as low complexity and homopolymer sequences
- Basecalling in and around methylation motifs
- De novo genome assembly quality
- Consensus accuracy (with and without trained polishing models)
- Short variant calling (SNPs and indels, with and without trained polishing models)
- Structural variants
If the validation meets the minimum criteria and the new model is an improvement on the currently-released models, it is then included in Oxford Nanopore's production software.
3. Live analysis
Introduction to live basecalling in MinKNOW
For MinION and PromethION 2 Solo, the MinKNOW software presents an option to basecall reads on the local computer. The basecalling is carried out live, as the read files are generated during a sequencing experiment.
Basecalling results are displayed in real-time in the MinKNOW user interface, and data is written out in the BAM or FASTQ file format.

Live alignment in MinKNOW
Basecalled reads can be aligned to a reference during the sequencing run. To do this, you will need to upload a reference FASTA or MMI file during run set-up, and optionally a BED file when there is a specific interest in a particular region of the reference (e.g. specific gene in a chromosome).
A reference file can contain multiple entries in the same file (e.g. multiple chromosomes), and alignment hits from these files are used to populate the alignment graphs which can be viewed on the MinKNOW UI. Alignment hits from BED files will appear in the sequencing TXT file generated in the data folder.
4. On-demand basecalling using the Dorado software
Dorado basecalling software
Dorado is a data processing toolkit that contains Oxford Nanopore basecalling algorithms, and several bioinformatic post-processing features. It is run from the command line in Windows, macOS, and on multiple Linux platforms. A selection of configuration files allow basecalling of DNA and RNA libraries prepared with current sequencing kits from Oxford Nanopore, and run on MinION or PromethION Flow Cells.
Further information about the Dorado basecaller can be found in the Dorado documentation.
Dorado availability
The Dorado basecalling software is available free of charge to the Nanopore Community and on GitHub. More details on installing and running the software are found in the Dorado GitHub repository and the Getting Started guide.
5. Basecall accuracy
Introduction to nanopore sequencing accuracy
Oxford Nanopore sequencing accuracy is shown as one of several metrics:
- per base quality score - denoted by the Phred Q-score
- raw read quality - this is calculated as an average from the q-scores, and this average quality is calibrated against accuracy.
- raw read accuracy - accuracy is calculated from an alignment to a reference sequence and counts insertions, deletions and substitutions as errors (bases correct / (insertions + deletions + bases aligned)), e.g. 99% accuracy can be interpreted as 99 out of 100 bases in a read were called correctly. Note that Oxford Nanopore represents the average raw read accuracy as the modal per read accuracy from a sequencing run.
Sequencing accuracy is determined by multiple factors, such as pore chemistry and basecalling algorithms. Improvements in these areas over the last several years have led to a steady increase in both single-molecule and consensus accuracy.
For more information about our latest accuracy data, please see the Accuracy page on the Oxford Nanopore website.
6. Barcoding options
Barcode design
Oxford Nanopore barcoding kits can place barcodes at the beginning and for some kits, also at the end of the strands for multiplexing several different samples in one sequencing experiment. The barcodes will reside in a kit-specific context sequence, and different kits will have different lengths of sequence before and after the barcode. However, the sequences of the barcodes themselves are identical, regardless of kit.
The regions of a barcode
A complete barcode arrangement comprises three sections:
- The upstream flanking region, which comes between the barcode and the sequencing adapter.
- The barcode sequence.
- The downstream flanking region, which comes between the barcode and the sample sequence.
The barcode sequences remain constant across almost all of Oxford Nanopore kits. For example, the flanking regions for barcode 10 in the Rapid Barcoding Kit (SQK-RBK114.24) are different from the flanking regions for barcode 10 in the Rapid PCR Barcoding Kit (SQK-RPB114.24), but the barcode sequence itself is the same. The exception is Native Barcoding kits, where the barcodes are the reverse complement of the standard barcodes.
Barcode and barcode flanking sequences can be found in the Chemistry technical document.
Barcode demultiplexing options
After a barcoded sequencing run has completed, the reads can be split into folders by barcode, using one of Oxford Nanopore's demultiplexing tools:
- Real-time barcode demultiplexing in MinKNOW
- Post-run barcode demultiplexing in MinKNOW
- Barcode demultiplexing in the Dorado basecall server
A brief description of the options is provided below.
Barcode demultiplexing in MinKNOW
MinKNOW currently uses Dorado for both basecalling and barcode demultiplexing. It performs barcode demultiplexing in real-time, as the sequencing run progresses. MinKNOW demultiplexing is also available as a post-run analysis option.
Barcode demultiplexing in Dorado
The barcoding algorithm in the Dorado basecall server uses a modified Needleman-Wunsch method. Each barcode is aligned to a section of the basecall, with a score assigned to each base in the sequence depending on whether the base was a match, mis-match or a gap. The combined scores for each barcode alignment are compared, and the barcode with the highest score is chosen as long as the score is above the defined threshold. The barcode sequences can be trimmed from the reads, as a command-line option.
7. File formats
For information about output file formats, please refer to Oxford Nanopore Output Specifications.
POD5 output
POD5 is an Oxford Nanopore-developed file format which stores nanopore data in an accessible way and replaces the legacy FAST5 format. This output also reads and writes data faster, uses less compute and has smaller raw data file size than FAST5.
For more information about the POD5 schema and contents, refer to POD5 file format.
Default read file location
Windows
C:\data\
macOS
/Library/MinKNOW/
Linux
/var/lib/MinKNOW/
Intermediate folder
The files in the intermediate folder store unprocessed raw signal data. Once raw signal processing is complete, POD5 files are generated and stored in the tmp folder, where local basecalling can proceed. These files are removed as processing proceeds or at the end of the run.
If the system encounters an issue, such as running out of space, the unprocessed data will not be cleared and will remain in the intermediate folder. Due to the real-time streaming nature of the system, this data cannot be processed after the run is stopped.
FASTQ output
FASTQ files are text files that contain sequence data for each read, and associated per-base quality scores. FASTQ files can be generated in MinKNOW and Dorado. The default is to write out 4000 reads per FASTQ file, although this number is configurable.
A single read sequence in a FASTQ file is described in four lines:
- Line 1 begins with a '@' and is followed by a header containing information about the sequencing run.
- Line 2 is the basecalled sequence (using A, C, T, G and N).
- Line 3 contains a '+'.
- Line 4 encodes the per-base quality scores for the sequence in Line 2.
BAM output
BAM files are output by MinKNOW and stand-alone Dorado software if alignment has been performed on the basecalled dataset. BAM files are also output when using the modified base models in MinKNOW and Dorado.
8. Data analysis in EPI2ME
The EPI2ME Desktop
The EPI2ME Desktop simplifies genomic data analysis for scientists without the need for bioinformatics expertise. With its intuitive interface, users can navigate through a collection of preconfigured workflows for best-practice genomic analyses of nanopore data.
EPI2ME is compatible with Windows, macOS or Linux and can be installed on a laptop, desktop computer, cluster or cloud service, as well as directly on Oxford Nanopore devices with computing capability (GridION, PromethION 2 Integrated, or PromethION 24). It can run locally or in the cloud and includes workflows running in real-time.
The EPI2ME platform uses the latest, internally validated, open-source analysis pipelines to deliver a growing range of streamlined, best-practice analysis workflows. Available EPI2ME workflows are listed here and include:
- Human genomics: All-in-one variant detection, including SNPs, SVs, CNVs, STRs, and methylation.
- Cancer genomics: Somatic variation detection from paired tumour/normal data.
- Single cell and transcriptomics: Comprehensive analysis of full-length transcripts.
- Microbiology and infectious disease: Real-time metagenomic species identification, and pathogen analysis workflows.
- Genome assembly: Plasmid and bacterial genome assembly and annotation.
- Targeted sequencing: Variant calling in amplicon sequences.
The workflows deliver intuitive and interactive reports and standard output files, including variants and methylation analysis.
In addition, EPI2ME provides industry-standard output files (e.g. VCF) as well as offering direct integration with selected third party analysis tools — enabling more comprehensive downstream data analysis.
EPI2ME workflows
EPI2ME workflows are developed using the Nextflow framework and, apart from being available through the graphical user interface of the EPI2ME Desktop, they are also available as open-source, command-line tools, downloadable from GitHub.
This framework facilitates the integration of community-developed Nextflow-based pipelines, which can be effortlessly imported into the user-friendly EPI2ME Desktop. It enables bioinformaticians to share their pipelines seamlessly with scientific colleagues, collaborators, and the broader community.
End-to-end workflows
EPI2ME is fully integrated into Oxford Nanopore’s end-to-end sequencing workflows, which have been developed to deliver optimal results for several common research applications. Detailed protocols guide you, step-by-step, through the complete sequencing process — from sample extraction recommendations through to sequencing and data analysis. Explore the full range of end-to-end application workflows.
9. Oxford Nanopore tools and pipelines
Oxford Nanopore GitHub repository contains a number of data analysis tools created by our R&D division. Most of the tools require some bioinformatics knowledge and use of the command line. Examples of software that are presented through this GitHub resource include experimental basecallers (Dorado, Remora), Modkit for the refinement of base modification results, and Medaka for the polishing of consensus sequence and calling of haploid variants.






