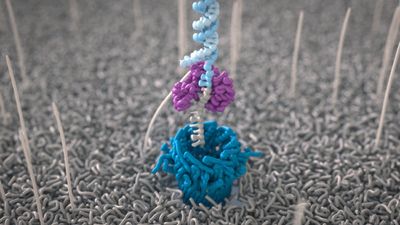
How Oxford Nanopore sequencing works
Oxford Nanopore has developed a new generation of DNA/RNA sequencing technology. It is the only sequencing technology that offers real-time analysis (for rapid insights), in fully scalable formats from pocket to population scale, that can analyse native DNA or RNA and sequence any length of fragment to achieve short to ultra-long read lengths.
Real-time
The only sequencing technology that offers real-time analysis
Fully scalable
From pocket to population-scale devices
Direct DNA/RNA sequencing
Call base modifications simultaneously with nucleotide sequence
Ultra-long reads
Improve assembly and characterise repetitive and difficult regions with ultra-long reads
Sequence anywhere
MinION is being used outside the traditional lab environment — taking the analysis to the sample.
Nanopores
All Oxford Nanopore sequencing devices use flow cells which contain an array of tiny holes — nanopores — embedded in an electro-resistant membrane. Each nanopore corresponds to its own electrode connected to a channel and sensor chip, which measures the electric current that flows through the nanopore. When a molecule passes through a nanopore, the current is disrupted to produce a characteristic ‘squiggle’. The squiggle is then decoded using basecalling algorithms to determine the DNA or RNA sequence in real time.
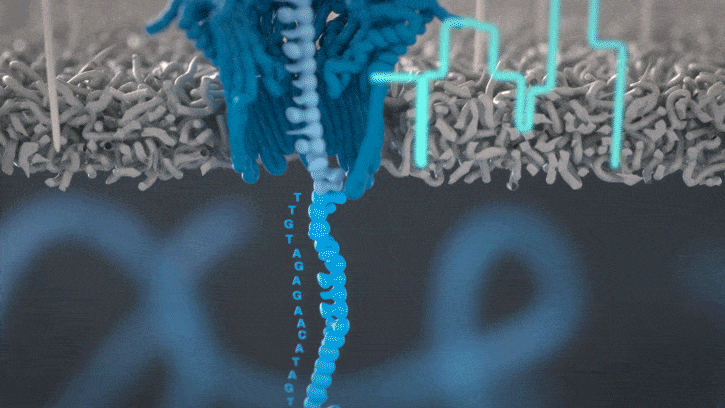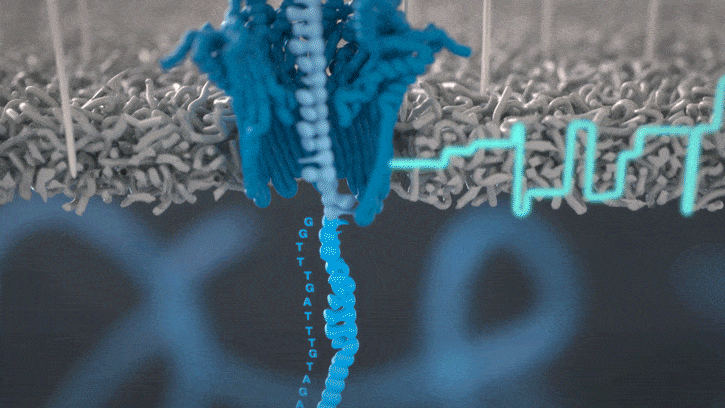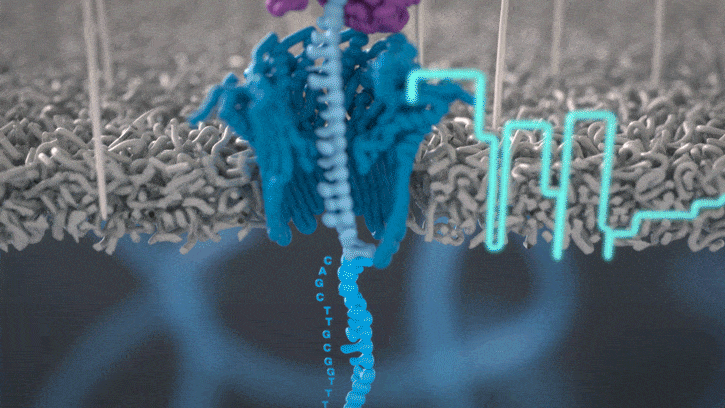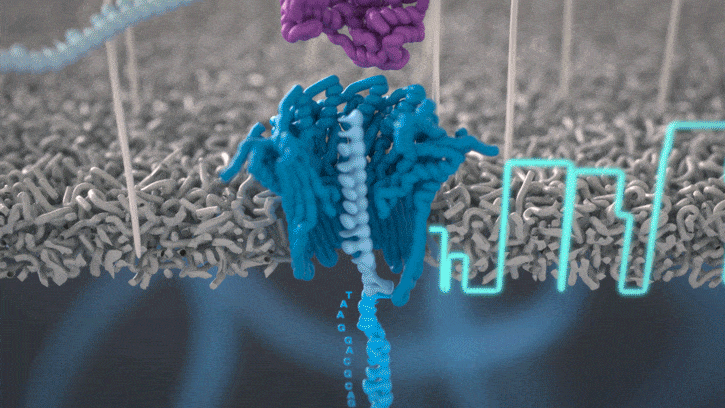
You can think of the current as water flowing through a pipe. When an object enters the pipe, the flow of water is disrupted, just as DNA disrupts the current as it passes through the nanopore.
DNA / RNA sequencing
A strand of DNA or RNA is made up of a sequence of different combinations of four nucleotide bases: A, T (or U for RNA), G, and C. Each base that passes through the nanopore can be identified through the characteristic disruption it causes to the current in real-time. This makes Oxford Nanopore sequencing unique, in that it is the only sequencing technology that enables direct, real-time analysis of short to ultra-long fragments of DNA/RNA, in fully scalable formats. Advantages of real-time sequencing include rapid access to time critical information (e.g. pathogen identification), the generation of early sample insights and more control over the sequencing experiment.
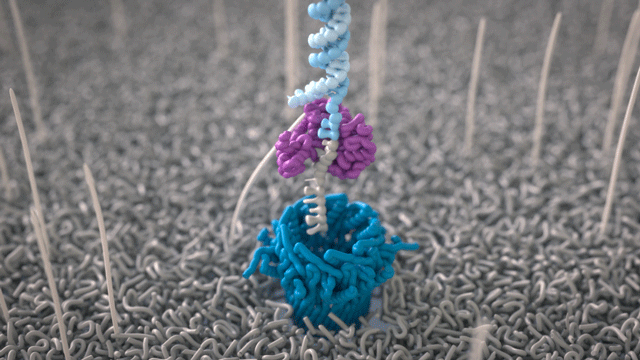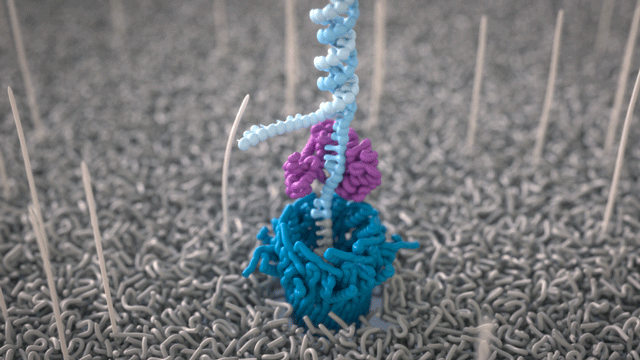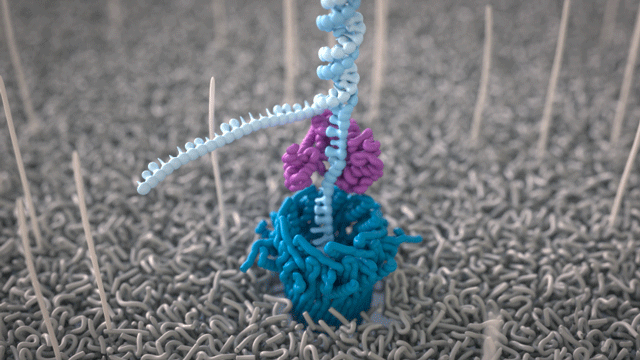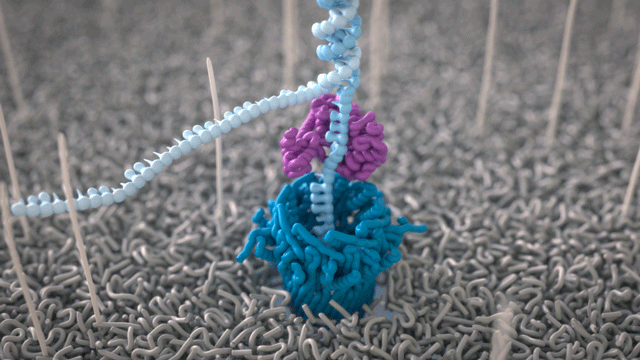
Why DNA / RNA?
DNA and RNA are molecules that are present in all living things. DNA is the genetic code of life, the instructions for building and operating an organism. RNA is primarily a messenger molecule, carrying instructions from the DNA code to control the synthesis of proteins — the building blocks of organisms. Sequencing can answer a range of biological questions, providing information on pathogen identity, genetic disease risk, or how an organism has evolved.

You can think of sequencing DNA/RNA likte reading music — the bases being the notes, which when played in he correct order, produces a recognisable the song. Oxford Nanopore sequencing is used to determine the sequence of DNA/RNA bases.
The power of long reads
Traditional methods are only able to sequence short lengths of DNA which must then be reassembled. It is therefore difficult to sequence repetitive regions for accurate genome assemblies without gaps, resolve large structural variations, or differentiate isoforms. Oxford Nanopore sequencing is limited only by the length of the DNA/RNA fragment presented to the pore and can therefore span entire repetitive regions, resolve structural variants, and differentiate between different isoforms. The ability to sequence native DNA and RNA without the requirement for amplification, eliminates PCR bias and allows for the identification of base modifications, such as methylation, alongside nucleotide sequence.
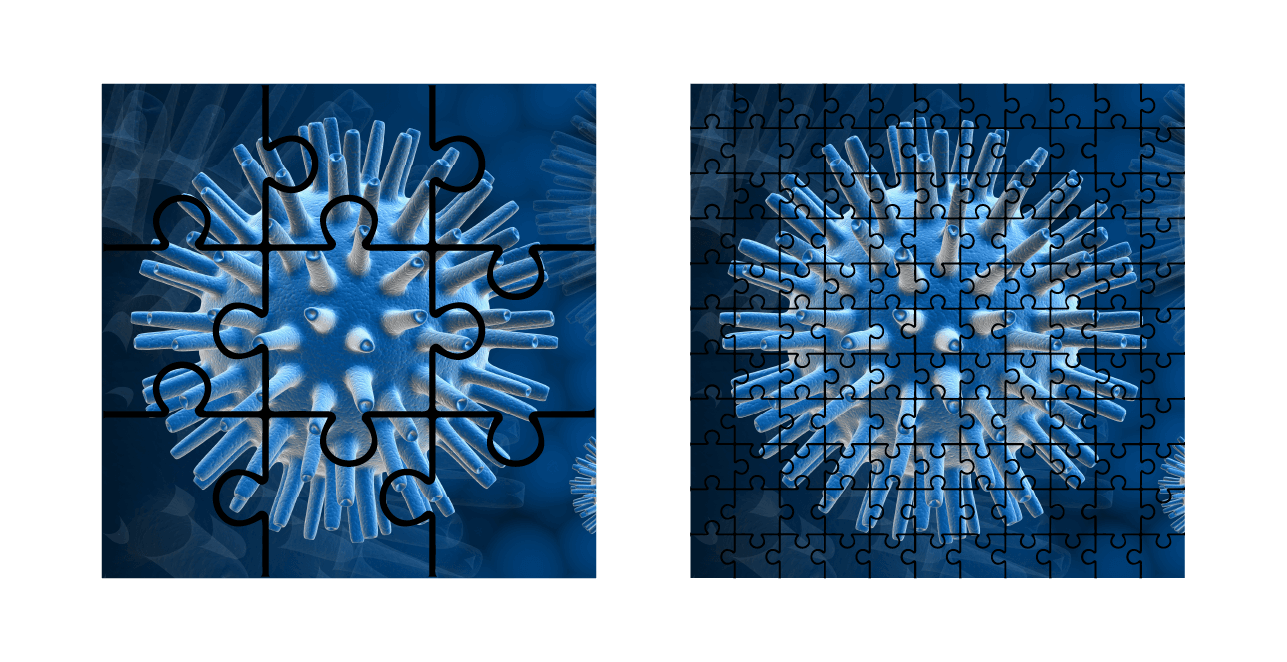

You can think of this like trying to complete two jigsaws of the same photograph — one with significantly larger pieces than the other. A jigsaw with only 9 pieces is much easier to assemble than one with 900.
Oxford Nanopore sequencing is fully scalable
Oxford Nanopore sequencing is the only sequencing technology to enable real-time analysis in fully scalable formats. From the pocket-sized MinION to the high-throughput, population-scale PromethION — scale nanopore sequencing to suit any experimental needs.
Get in touch
Subscribe
Talk to us
If you have any questions about our products or services, chat directly with a member of our sales team.
Book a sales call
To book a call with one of our sales team, please click below.











